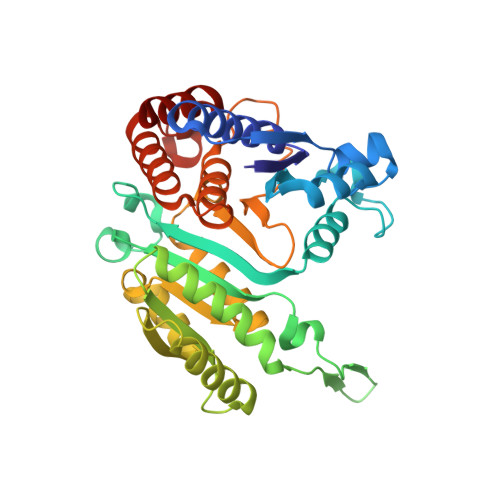
Structure of 3-isopropylmalate dehydrogenase in complex with NAD+: ligand-induced loop closing and mechanism for cofactor specificity.
Hurley, J.H., Dean, A.M.(1994) Structure 2: 1007-1016
- PubMed: 7881901 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(94)00104-9
- Primary Citation Related Structures:
1HEX - PubMed Abstract:
The leucine biosynthetic enzyme 3-isopropylmalate dehydrogenase (IMDH) belongs to a unique class of bifunctional decarboxylating dehydrogenases. The two best-known members of this family, IMDH and isocitrate dehydrogenase (IDH), share a common structural framework and catalytic mechanism but have different substrate and cofactor specificities. IMDH is NAD(+)-dependent, while IDHs occur in both NAD(+)-dependent and NADP(+)-dependent forms. We have co-crystallized Thermus thermophilus IMDH with NAD+ and have determined the structure at 2.5 A resolution. NAD+ binds in an extended conformation. Comparisons with the structure in the absence of cofactor show that binding induces structural changes of up to 2.5 A in the five loops which form the dinucleotide-binding site. The adenine and diphosphate moieties of NAD+ are bound via interactions which are also present in the NADP(+)-IDH complex. Amino acids which interact with the NADP+ 2'-phosphate in IDH are substituted or absent in IMDH. The adenosine ribose forms two hydrogen bonds with Asp278, and the nicotinamide and nicotinamide ribose interact with Glu87 and Asp78, all unique to IMDH. NAD+ binding induces a conformational transition in IMDH, resulting in a structure that is intermediate between the most 'open' and 'closed' decarboxylating dehydrogenase conformations. Physiological specificity of IMDH for NAD+ versus NADP+ can be explained by the unique interaction between Asp278 and the free 2'-hydroxyl of the NAD+ adenosine, discrimination against the presence of the 2'-phosphate by the negative charge on Asp278, and the absence of potential favorable interactions with the 2'-phosphate of NADP+.
- Laboratory of Molecular Biology, National Institute of Diabetes, Digestive and Kidney Diseases, National Institutes of Health, Bethesda, MD 20892-0580.
Organizational Affiliation: