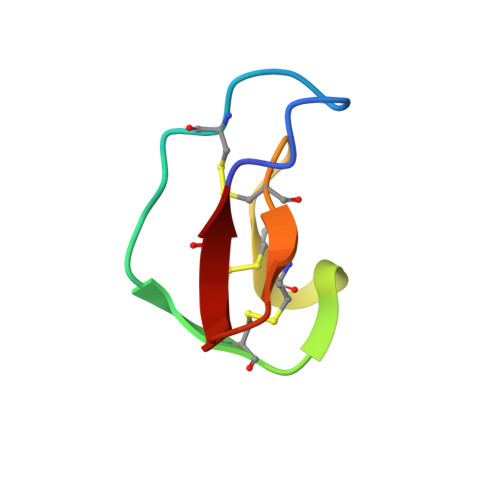
Solution Structure of the Squash Trypsin Inhibitor Mcoti-II. A New Family for Cyclic Knottins
Heitz, A., Hernandez, J.-F., Gagnon, J., Hong, T.T., Pham, T.T.C., Nguyen, T.M., Le-Nguyen, D., Chiche, L.(2001) Biochemistry 40: 7973
- PubMed: 11434766 Search on PubMed
- DOI: https://doi.org/10.1021/bi0106639
- Primary Citation Related Structures:
1HA9 - PubMed Abstract:
The "knottin" fold is a stable cysteine-rich scaffold, in which one disulfide crosses the macrocycle made by two other disulfides and the connecting backbone segments. This scaffold is found in several protein families with no evolutionary relationships. In the past few years, several homologous peptides from the Rubiaceae and Violaceae families were shown to define a new structural family based on macrocyclic knottin fold. We recently isolated from Momordica cochinchinensis seeds the first known macrocyclic squash trypsin inhibitors. These compounds are the first members of a new family of cyclic knottins. In this paper, we present NMR structural studies of one of them, MCoTI-II, and of a beta-Asp rearranged form, MCoTI-IIb. Both compounds display similar and well-defined conformations. These cyclic squash inhibitors share a similar conformation with noncyclic squash inhibitors such as CPTI-II, and it is postulated that the main effect of the cyclization is a reduced sensitivity to exo-proteases. On the contrary, clear differences were detected with the three-dimensional structures of other known cyclic knottins, i.e., kalata B1 or circulin A. The two-disulfide cystine-stabilized beta-sheet motif [Heitz et al. (1999) Biochemistry 38, 10615-10625] is conserved in the two families, whereas in the C-to-N linker, one disulfide bridge and one loop are differently located. The molecular surface of MCoTI-II is almost entirely charged in contrast to circulin A that displays a well-marked amphiphilic character. These differences might explain why the isolated macrocyclic squash inhibitors from M. cochinchinensis display no significant antibacterial activity, whereas circulins and kalata B1 do.
- Centre de Biochimie Structurale, UMR5048 CNRS-Université Montpellier I, UMR554 INSERM-Université Montpellier I, Faculté de pharmacie, 15 avenue Charles Flahault, 34060 Montpellier, France.
Organizational Affiliation: