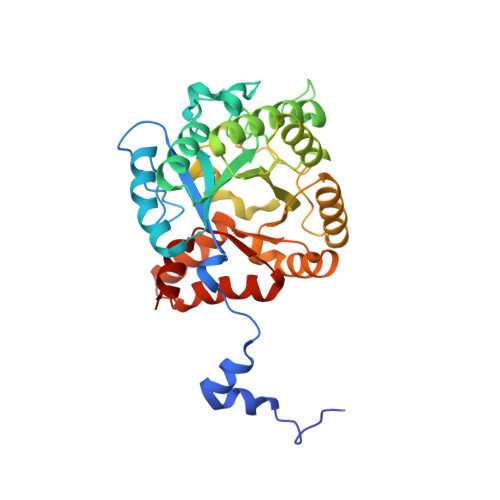
Structure of porphobilinogen synthase from Pseudomonas aeruginosa in complex with 5-fluorolevulinic acid suggests a double Schiff base mechanism.
Frere, F., Schubert, W.D., Stauffer, F., Frankenberg, N., Neier, R., Jahn, D., Heinz, D.W.(2002) J Mol Biology 320: 237-247
- PubMed: 12079382 Search on PubMed
- DOI: https://doi.org/10.1016/S0022-2836(02)00472-2
- Primary Citation Related Structures:
1GZG - PubMed Abstract:
All natural tetrapyrroles, including hemes, chlorophylls and vitamin B12, share porphobilinogen (PBG) as a common precursor. Porphobilinogen synthase (PBGS) synthesizes PBG through the asymmetric condensation of two molecules of aminolevulinic acid (ALA). Crystal structures of PBGS from various sources confirm the presence of two distinct binding sites for each ALA molecule, termed A and P. We have solved the structure of the active-site variant D139N of the Mg2+-dependent PBGS from Pseudomonas aeruginosa in complex with the inhibitor 5-fluorolevulinic acid at high resolution. Uniquely, full occupancy of both substrate binding sites each by a single substrate-like molecule was observed. Both inhibitor molecules are covalently bound to two conserved, active-site lysine residues, Lys205 and Lys260, through Schiff bases. The active site now also contains a monovalent cation that may critically enhance enzymatic activity. Based on these structural data, we postulate a catalytic mechanism for P. aeruginosa PBGS initiated by a C-C bond formation between A and P-side ALA, followed by the formation of the intersubstrate Schiff base yielding the product PBG.
- Institute of Microbiology, Technical University Braunschweig, Spielmannstrasse 7, D-38106 Braunschweig, Germany.
Organizational Affiliation: