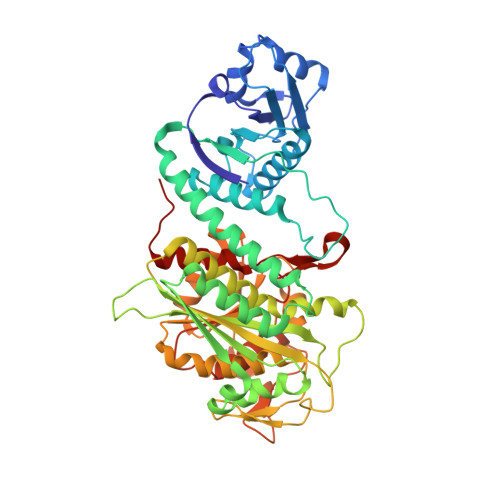
X-Ray Structure of Aminopeptidase a from Escherichia Coli and a Model for the Nucleoprotein Complex in Xer Site-Specific Recombination
Straeter, N., Sherratt, D.J., Colloms, S.D.(1999) EMBO J 18: 4513
- PubMed: 10449417 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/18.16.4513
- Primary Citation Related Structures:
1GYT - PubMed Abstract:
The structure of aminopeptidase A (PepA), which functions as a DNA-binding protein in Xer site-specific recombination and in transcriptional control of the carAB operon in Escherichia coli, has been determined at 2.5 A resolution. In Xer recombination at cer, PepA and the arginine repressor (ArgR) serve as accessory proteins, ensuring that recombination is exclusively intramolecular. In contrast, PepA homologues from other species have no known DNA-binding activity and are not implicated in transcriptional regulation or control of site-specific recombination. PepA comprises two domains, which have similar folds to the two domains of bovine lens leucine aminopeptidase (LAP). However, the N-terminal domain of PepA, which probably plays a significant role in DNA binding, is rotated by 19 degrees compared with its position in LAP. PepA is a homohexamer of 32 symmetry. A groove that runs from one trimer face across the 2-fold molecular axis to the other trimer face is proposed to be the DNA-binding site. Molecular modelling supports a structure of the Xer complex in which PepA, ArgR and a second PepA molecule are sandwiched along their 3-fold molecular axes, and the accessory sequences of the two recombination sites wrap around the accessory proteins as a right-handed superhelix such that three negative supercoils are trapped.
- Institut für Kristallographie, Freie Universität Berlin, Takustrasse 6, 14195 Berlin, Germany. strater@chemie.fu-berlin.de
Organizational Affiliation: