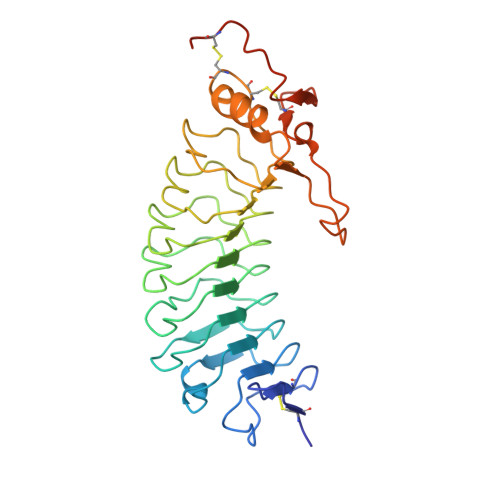
Crystal Structure of the Platelet Glycoprotein Ib-Alpha N-Terminal Domain Reveals an Unmasking Mechanism of Receptor Activation
Uff, S., Clemetson, J.M., Harrison, T., Clemetson, K.J., Emsley, J.(2002) J Biological Chem 277: 35657
- PubMed: 12087105 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M205271200
- Primary Citation Related Structures:
1GWB - PubMed Abstract:
Glycoprotein Ib (GPIb) is a platelet receptor with a critical role in mediating the arrest of platelets at sites of vascular damage. GPIb binds to the A1 domain of von Willebrand factor (vWF-A1) at high blood shear, initiating platelet adhesion and contributing to the formation of a thrombus. To investigate the molecular basis of GPIb regulation and ligand binding, we have determined the structure of the N-terminal domain of the GPIb(alpha) chain (residues 1-279). This structure is the first determined from the cell adhesion/signaling class of leucine-rich repeat (LRR) proteins and reveals the topology of the characteristic disulfide-bonded flanking regions. The fold consists of an N-terminal beta-hairpin, eight leucine-rich repeats, a disulfide-bonded loop, and a C-terminal anionic region. The structure also demonstrates a novel LRR motif in the form of an M-shaped arrangement of three tandem beta-turns. Negatively charged binding surfaces on the LRR concave face and anionic region indicate two-step binding kinetics to vWF-A1, which can be regulated by an unmasking mechanism involving conformational change of a key loop. Using molecular docking of the GPIb and vWF-A1 crystal structures, we were also able to model the GPIb.vWF-A1 complex.
- Department of Biochemistry, University of Leicester, Leicester LE1 7RH, United Kingdom.
Organizational Affiliation: