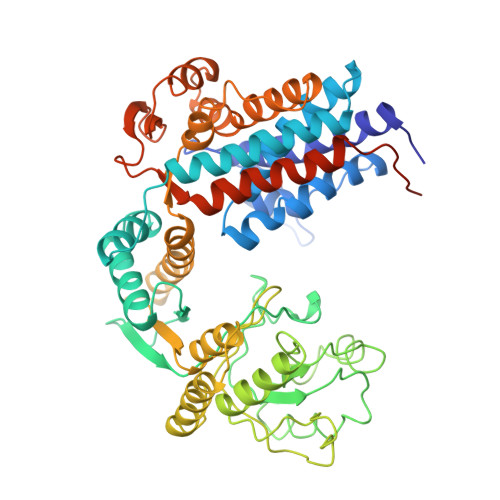
The crystal structure of the bacterial chaperonin GroEL at 2.8 A.
Braig, K., Otwinowski, Z., Hegde, R., Boisvert, D.C., Joachimiak, A., Horwich, A.L., Sigler, P.B.(1994) Nature 371: 578-586
- PubMed: 7935790
- DOI: https://doi.org/10.1038/371578a0
- Primary Citation Related Structures:
1GRL - PubMed Abstract:
The crystal structure of Escherichia coli GroEL shows a porous cylinder of 14 subunits made of two nearly 7-fold rotationally symmetrical rings stacked back-to-back with dyad symmetry. The subunits consist of three domains: a large equatorial domain that forms the foundation of the assembly at its waist and holds the rings together; a large loosely structured apical domain that forms the ends of the cylinder; and a small slender intermediate domain that connects the two, creating side windows. The three-dimensional structure places most of the mutationally defined functional sites on the channel walls and its outward invaginations, and at the ends of the cylinder.
- Department of Genetics, Yale University School of Medicine, Boyer Center, New Haven, Connecticut 06510.
Organizational Affiliation: