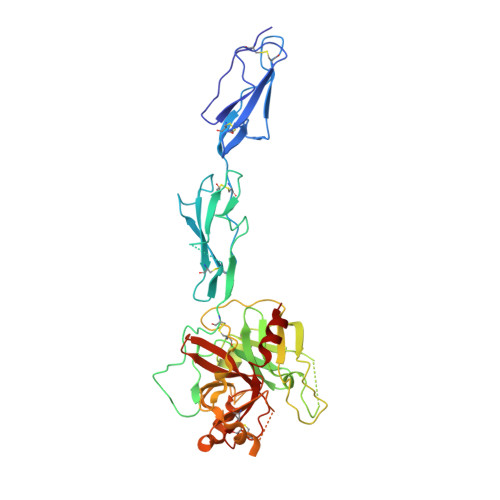
The Crystal Structure of the Zymogen Catalytic Domain of Complement Protease C1R Reveals that a Disruptive Mechanical Stress is Required to Trigger Activation of the C1 Complex.
Budayova-Spano, M., Lacroix, M., Thielens, N., Arlaud, G., Fontecilla-Camps, J.C., Gaboriaud, C.(2002) EMBO J 21: 231
- PubMed: 11823416 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/21.3.231
- Primary Citation Related Structures:
1GPZ - PubMed Abstract:
C1r is the modular serine protease (SP) that mediates autolytic activation of C1, the macromolecular complex that triggers the classical pathway of complement. The crystal structure of a mutated, proenzyme form of the catalytic domain of human C1r, comprising the first and second complement control protein modules (CCP1, CCP2) and the SP domain has been solved and refined to 2.9 A resolution. The domain associates as a homodimer with an elongated head-to-tail structure featuring a central opening and involving interactions between the CCP1 module of one monomer and the SP domain of its counterpart. Consequently, the catalytic site of one monomer and the cleavage site of the other are located at opposite ends of the dimer. The structure reveals unusual features in the SP domain and provides strong support for the hypothesis that C1r activation in C1 is triggered by a mechanical stress caused by target recognition that disrupts the CCP1-SP interfaces and allows formation of transient states involving important conformational changes.
- Laboratoire de Cristallographie et Cristollogénèse des Protéines, Institut de Biologie Structurale Jean-Pierre Ebel, CEA-CNRS-UJF, 41 rue Jules Horowitz, F-38027 Grenoble cedex 1, France.
Organizational Affiliation: