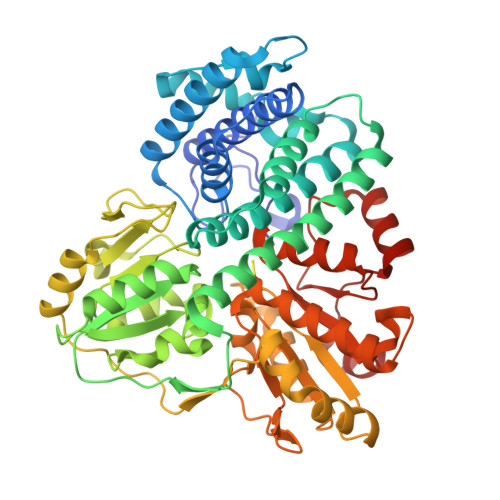
Hybrid Cluster Proteins (Hcps) from Desulfovibrio Desulfuricans Atcc 27774 and Desulfovibrio Vulgaris (Hildenborough): X-Ray Structures at 1.25 A Resolution Using Synchrotron Radiation.
Macedo, S., Mitchell, E.P., Romao, C.V., Cooper, S.J., Coelho, R., Liu, M.Y., Xavier, A.V., Legall, J., Bailey, S., Garner, D.C., Hagen, W.R., Teixeira, M., Carrondo, M.A., Lindley, P.(2002) J Biol Inorg Chem 7: 514
- PubMed: 11941509 Search on PubMed
- DOI: https://doi.org/10.1007/s00775-001-0326-y
- Primary Citation Related Structures:
1GN9, 1GNL, 1GNT - PubMed Abstract:
The structures of the hybrid cluster proteins (HCPs) from the sulfate-reducing bacteria Desulfovibrio desulfuricans (ATCC 27774) and Desulfovibrio vulgaris (Hildenborough) have been elucidated at a resolution of 1.25 A using X-ray synchrotron radiation techniques. In the case of the D. desulfuricans protein, protein isolation, purification, crystallization and X-ray data collection were carried out under strict anaerobic conditions, whereas for the D. vulgaris protein the conditions were aerobic. However, both structures are essentially the same, comprising three domains and two iron-sulfur centres. One of these centres situated near the exterior of the molecules in domain 1 is a cubane [4Fe-4S] cluster, whereas the other, located at the interface of the three domains, contains the unusual four-iron cluster initially found in the D. vulgaris protein. Details of the structures and the associated EPR spectroscopy of the D. desulfuricans protein are reported herein. These structures show that the nature of the hybrid cluster, containing both oxygen and sulfur bridges, is independent of the presence of oxygen in the isolation and crystallization procedure and also does not vary significantly with changes in the oxidation state. The structures and amino acid sequences of the HCP are compared with the recently elucidated structure of the catalytic subunit of a carbon monoxide dehydrogenase from Carboxydothermus hydrogenoformans and related dehydrogenases. Electronic supplementary material to this paper can be obtained by using the Springer Link server located at http://dx.doi.org/10.1007/s00775-001-0326-y.
- Instituto de Tecnologia Química e Biológica, Universidade Nova de Lisboa, Av. República, EAN, Apartado 127, 2781-901 Oeiras, Portugal.
Organizational Affiliation: