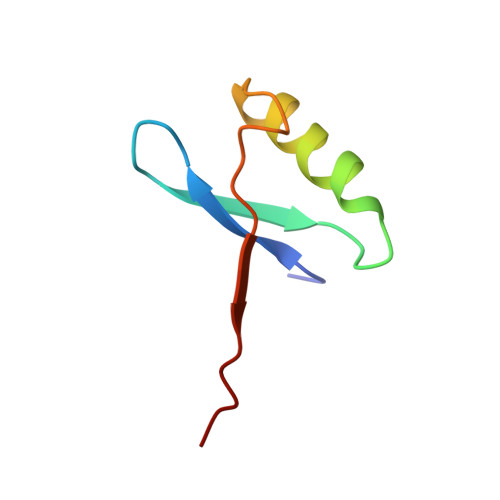
Structure and Properties of a Dimeric N-Terminal Fragment of Human Ubiquitin.
Bolton, D., Evans, P.A., Stott, K., Broadhurst, R.W.(2001) J Mol Biology 314: 773
- PubMed: 11733996 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2001.5181
- Primary Citation Related Structures:
1GJZ - PubMed Abstract:
Previous peptide dissection and kinetic experiments have indicated that in vitro folding of ubiquitin may proceed via transient species in which native-like structure has been acquired in the first 45 residues. A peptide fragment, UQ(1-51), encompassing residues 1 to 51 of ubiquitin was produced in order to test whether this portion has propensity for independent self-assembly. Surprisingly, the construct formed a folded symmetrical dimer that was stabilised by 0.8 M sodium sulphate at 298 K (the S state). The solution structure of the UQ(1-51) dimer was determined by multinuclear NMR spectroscopy. Each subunit of UQ(1-51) consists of an N-terminal beta-hairpin followed by an alpha-helix and a final beta-strand, with orientations similar to intact ubiquitin. The dimer is formed by the third beta-strand of one subunit interleaving between the hairpin and third strand of the other to give a six-stranded beta-sheet, with the two alpha-helices sitting on top. The helix-helix and strand portions of the dimer interface also mimic related features in the structure of ubiquitin. The structural specificity of the UQ(1-51) peptide is tuneable: as the concentration of sodium sulphate is decreased, near-native alternative conformations are populated in slow chemical exchange. Magnetization transfer experiments were performed to characterize the various species present in 0.35 M sodium sulphate, namely the S state and two minor forms. Chemical shift differences suggest that one minor form is very similar to the S state, while the other experiences a significant conformational change in the third strand. A segmental rearrangement of the third strand in one subunit of the S state would render the dimer asymmetric, accounting for most of our results. Similar small-scale transitions in proteins are often invoked to explain solvent exchange at backbone amide proton sites that have an intermediate level of protection.
- Cambridge Centre for Molecular Recognition Department of Biochemistry, University of Cambridge, 80 Tennis Court Road, Cambridge, CB2 1GA, UK. r.w.broadhurst@bio.cam.ac.uk
Organizational Affiliation: