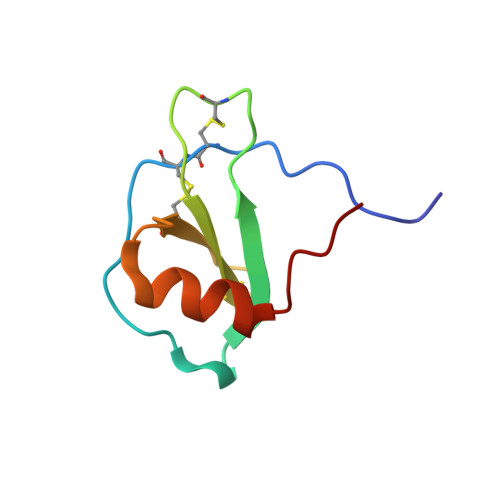
NMR solution structure and backbone dynamics of the CC chemokine eotaxin-3.
Ye, J., Mayer, K.L., Mayer, M.R., Stone, M.J.(2001) Biochemistry 40: 7820-7831
- PubMed: 11425309 Search on PubMed
- DOI: https://doi.org/10.1021/bi010252s
- Primary Citation Related Structures:
1G2S, 1G2T - PubMed Abstract:
Eotaxin-3 is one of three related chemokines that specifically activate chemokine receptor CCR3. We report the 3D structure and backbone dynamics of eotaxin-3 determined by NMR spectroscopy. Eotaxin-3 is monomeric under the conditions in this study and consists of an unstructured N-terminus before the first two conserved cysteine residues, an irregularly structured N-loop following the second conserved cysteine, a single turn of 3(10)-helix, a three-stranded antiparallel beta-sheet, an alpha-helix, and an unstructured C-terminal tail. As in other chemokines, the alpha-helix packs against one face of the beta-sheet. The average backbone and heavy atom rmsd values of the 20 structures (residues 9-65) are 0.44 and 1.01 A, respectively. A comparison between the structures of eotaxin-3 and related chemokines suggests that the electrostatic potential in the vicinity of a surface groove and the structure of the beta2-beta3 turn may be important for maintaining receptor specificity. The backbone dynamics of eotaxin-3 were determined from 15N NMR relaxation data using the extended model free dynamics formalism. Large amplitude motions on the picosecond to nanosecond time scale were observed in both termini and in some residues in the N-loop, the beta1-beta2 turn, and the beta3 strand; the location of these residues suggests a possible role for dynamics in receptor binding and activation. In contrast to eotaxin, eotaxin-3 exhibits no substantial mobility on the microsecond to millisecond time scale.
- Department of Chemistry, Indiana University, Bloomington, Indiana 47405-0001, USA.
Organizational Affiliation: