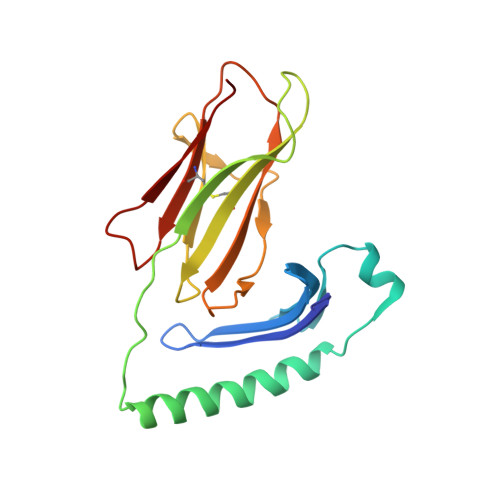
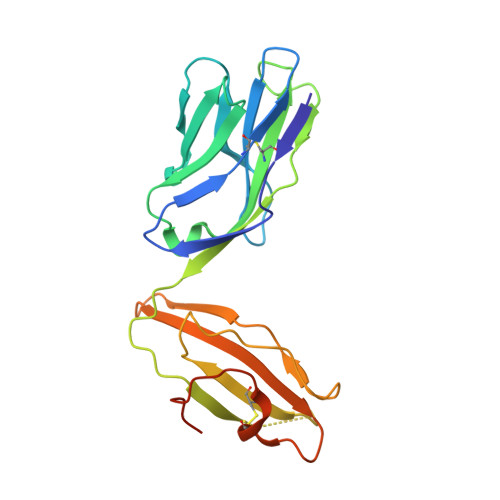
Structure of a covalently stabilized complex of a human alphabeta T-cell receptor, influenza HA peptide and MHC class II molecule, HLA-DR1.
Hennecke, J., Carfi, A., Wiley, D.C.(2000) EMBO J 19: 5611-5624
- PubMed: 11060013 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/emboj/19.21.5611
- Primary Citation Related Structures:
1FYT - PubMed Abstract:
An alphabeta T-cell receptor (alphabetaTCR)/hemagglutinin (HA) peptide/human leukocyte antigen (HLA)-DR1 complex was stabilized by flexibly linking the HA peptide with the human HA1.7 alphabetaTCR, to increase the local concentration of the interacting proteins once the peptide has been loaded onto the major histocompatibility complex (MHC) molecule. The structure of the complex, determined by X-ray crystallography, has a binding mode similar to that of the human B7 alphabetaTCR on a pMHCI molecule. Twelve of the 15 MHC residues contacted are at the same positions observed earlier in class I MHC/peptide/TCR complexes. One contact, to an MHC loop outside the peptide-binding site, is conserved and specific to pMHCII complexes. TCR gene usage in the response to HA/HLA-DR appears to conserve charged interactions between three lysines of the peptide and acidic residues on the TCR.
- Department of Molecular and Cellular Biology, Harvard University, Howard Hughes Medical Institute, 7 Divinity Avenue, Cambridge, MA 02138, USA.
Organizational Affiliation: