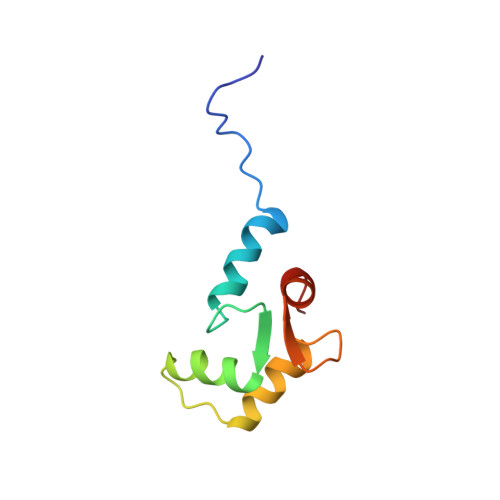
Solution structures of the C-terminal domain of cardiac troponin C free and bound to the N-terminal domain of cardiac troponin I.
Gasmi-Seabrook, G.M., Howarth, J.W., Finley, N., Abusamhadneh, E., Gaponenko, V., Brito, R.M., Solaro, R.J., Rosevear, P.R.(1999) Biochemistry 38: 8313-8322
- PubMed: 10387077
- DOI: https://doi.org/10.1021/bi9902642
- Primary Citation Related Structures:
1FI5 - PubMed Abstract:
The N-terminal domain of cardiac troponin I (cTnI) comprising residues 33-80 and lacking the cardiac-specific amino terminus forms a stable binary complex with the C-terminal domain of cardiac troponin C (cTnC) comprising residues 81-161. We have utilized heteronuclear multidimensional NMR to assign the backbone and side-chain resonances of Ca2+-saturated cTnC(81-161) both free and bound to cTnI(33-80). No significant differences were observed between secondary structural elements determined for free and cTnI(33-80)-bound cTnC(81-161). We have determined solution structures of Ca2+-saturated cTnC(81-161) free and bound to cTnI(33-80). While the tertiary structure of cTnC(81-161) is qualitatively similar to that observed free in solution, the binding of cTnI(33-80) results mainly in an opening of the structure and movement of the loop region between helices F and G. Together, these movements provide the binding site for the N-terminal domain of cTnI. The putative binding site for cTnI(33-80) was determined by mapping amide proton and nitrogen chemical shift changes, induced by the binding of cTnI(33-80), onto the C-terminal cTnC structure. The binding interface for cTnI(33-80), as suggested from chemical shift changes, involves predominantly hydrophobic interactions located in the expanded hydrophobic pocket. The largest chemical shift changes were observed in the loop region connecting helices F and G. Inspection of available TnC sequences reveals that these residues are highly conserved, suggesting a common binding motif for the Ca2+/Mg2+-dependent interaction site in the TnC/TnI complex.
- Department of Molecular Genetics, University of Cincinnati, College of Medicine, Ohio 45267, USA.
Organizational Affiliation: