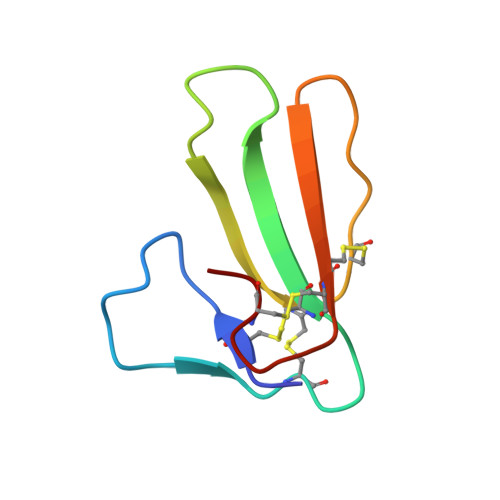
1.9-A resolution structure of fasciculin 1, an anti-acetylcholinesterase toxin from green mamba snake venom.
le Du, M.H., Marchot, P., Bougis, P.E., Fontecilla-Camps, J.C.(1992) J Biological Chem 267: 22122-22130
- PubMed: 1429564 Search on PubMed
- DOI: https://doi.org/10.2210/pdb1fas/pdb
- Primary Citation Related Structures:
1FAS - PubMed Abstract:
The crystal structure of fasciculin 1, a potent acetylcholinesterase inhibitor from green mamba snake venom, has been solved by the multiple isomorphous replacement method complemented with anomalous scattering and subsequently refined at 1.9-A resolution. The overall structure of fasciculin is similar to those of the short alpha-neurotoxins and cardiotoxins, with a dense core rich in disulfide bridges and three long loops disposed as the central fingers of a hand. A comparison of these three prototypic toxin types shows that fasciculin 1 has structural features that are intermediate between those of the other two molecules. Its core region, which can be defined as a continuous stretch of conserved residues, is very similar to that of erabutoxin b, whereas the orientation of its long loops resembles that of cardiotoxin VII4. This result introduces a new element in the study of phylogenetic relationships of snake toxins and suggests that, after divergency from an ancestral gene, convergent evolution may have played an important factor in the evolution of these proteins. In fasciculin 1, several arginine and lysine residues are well ordered and relatively exposed to the solvent medium and may play a role in the binding to the peripheral site of acetylcholinesterases.
- Laboratoire de Cristallographie et Cristallogénèse des Protéines, Département d'Ingéniérie et d'Etudes des Protéines, DSV, CENG, Grenoble, France.
Organizational Affiliation: