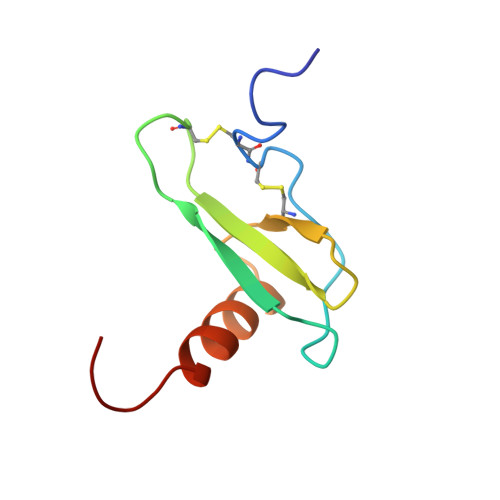
Solution structure of eotaxin, a chemokine that selectively recruits eosinophils in allergic inflammation.
Crump, M.P., Rajarathnam, K., Kim, K.S., Clark-Lewis, I., Sykes, B.D.(1998) J Biological Chem 273: 22471-22479
- PubMed: 9712872 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.273.35.22471
- Primary Citation Related Structures:
1EOT, 2EOT - PubMed Abstract:
The solution structure of the CCR3-specific chemokine, eotaxin, has been determined by NMR spectroscopy. The quaternary structure of eotaxin was investigated by ultracentrifugation and NMR, and it was found to be in equilibrium between monomer and dimer under a wide range of conditions. At pH
- Protein Engineering Network of Centres of Excellence (PENCE) and Department of Biochemistry, University of Alberta, Edmonton, Alberta, Canada T6G 2S2.
Organizational Affiliation: