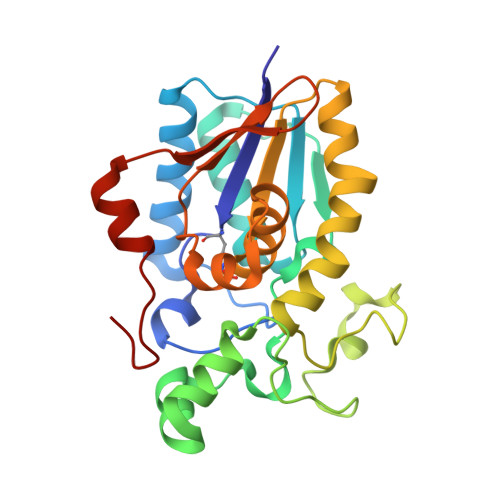
High Resolution Structure of the Phosphohistidine-Activated Form of Escherichia Coli Cofactor-Dependent Phosphoglycerate Mutase.
Bond, C.S., White, M.F., Hunter, W.N.(2001) J Biological Chem 276: 3247
- PubMed: 11038361 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M007318200
- Primary Citation Related Structures:
1E58 - PubMed Abstract:
The active conformation of the dimeric cofactor-dependent phosphoglycerate mutase (dPGM) from Escherichia coli has been elucidated by crystallographic methods to a resolution of 1.25 A (R-factor 0.121; R-free 0.168). The active site residue His(10), central in the catalytic mechanism of dPGM, is present as a phosphohistidine with occupancy of 0.28. The structural changes on histidine phosphorylation highlight various features that are significant in the catalytic mechanism. The C-terminal 10-residue tail, which is not observed in previous dPGM structures, is well ordered and interacts with residues implicated in substrate binding; the displacement of a loop adjacent to the active histidine brings previously overlooked residues into positions where they may directly influence catalysis. E. coli dPGM, like the mammalian dPGMs, is a dimer, whereas previous structural work has concentrated on monomeric and tetrameric yeast forms. We can now analyze the sequence differences that cause this variation of quaternary structure.
- Wellcome Trust Biocentre, University of Dundee, Dundee DD1 5EH, United Kingdom.
Organizational Affiliation: