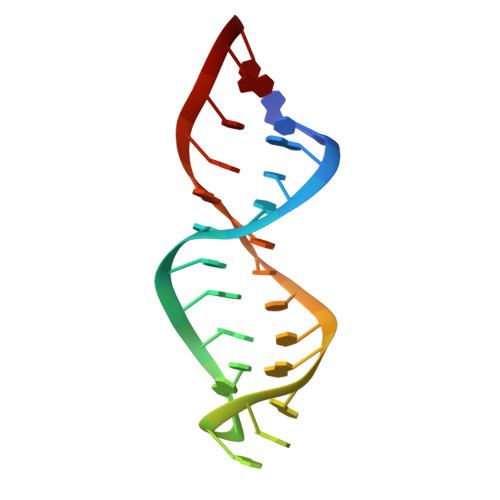
Structure of the Ribozyme Substrate Hairpin of Neurospora Vs RNA: A Close Look at the Cleavage Site
Michiels, P.J.A., Schouten, C.H.J., Hilbers, C.W., Heus, H.A.(2000) RNA 6: 1821
- PubMed: 11142381 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1017/s1355838200001394
- Primary Citation Related Structures:
1E4P - PubMed Abstract:
The cleavage site of the Neurospora VS RNA ribozyme is located in a separate hairpin domain containing a hexanucleotide internal loop with an A-C mismatch and two adjacent G-A mismatches. The solution structure of the internal loop and helix la of the ribozyme substrate hairpin has been determined by nuclear magnetic resonance (NMR) spectroscopy. The 2 nt in the internal loop, flanking the cleavage site, a guanine and adenine, are involved in two sheared G.A base pairs similar to the magnesium ion-binding site of the hammerhead ribozyme. Adjacent to the tandem G.A base pairs, the adenine and cytidine, which are important for cleavage, form a noncanonical wobble A+-C base pair. The dynamic properties of the internal loop and details of the high-resolution structure support the view that the hairpin structure represents a ground state, which has to undergo a conformational change prior to cleavage. Results of chemical modification and mutagenesis data of the Neurospora VS RNA ribozyme can be explained in context with the present three-dimensional structure.
- NSR Center for Molecular Structure, Design and Synthesis, Laboratory of Biophysical Chemistry, University of Nijmegen, The Netherlands.
Organizational Affiliation: