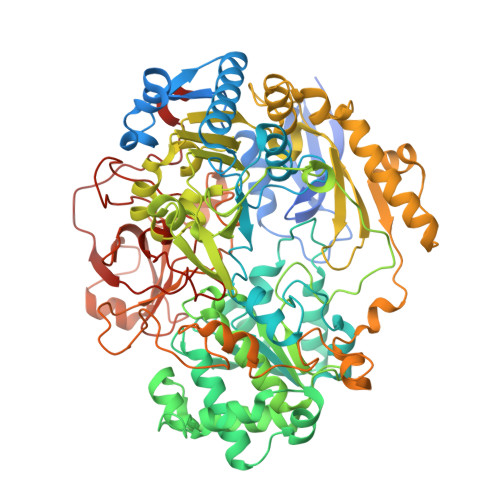
Dimethylsulfoxide Reductase: An Enzyme Capable of Catalysis with Either Molybdenum or Tungsten at the Active Site
Stewart, L.J., Bailey, S., Bennett, B., Charnock, J.M., Garner, C.D., Mcalpine, A.S.(2000) J Mol Biology 299: 593
- PubMed: 10835270 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.2000.3702
- Primary Citation Related Structures:
1E18 - PubMed Abstract:
DMSO reductase (DMSOR) from Rhodobacter capsulatus, well-characterised as a molybdoenzyme, will bind tungsten. Protein crystallography has shown that tungsten in W-DMSOR is ligated by the dithiolene group of the two pyranopterins, the oxygen atom of Ser147 plus another oxygen atom, and is located in a very similar site to that of molybdenum in Mo-DMSOR. These conclusions are consistent with W L(III)-edge X-ray absorption, EPR and UV/visible spectroscopic data. W-DMSOR is significantly more active than Mo-DMSOR in catalysing the reduction of DMSO but, in contrast to the latter, shows no significant ability to catalyse the oxidation of DMS.
- CLRC Daresbury Laboratory, Daresbury, Warrington, Cheshire, WA4 4AD, UK.
Organizational Affiliation: