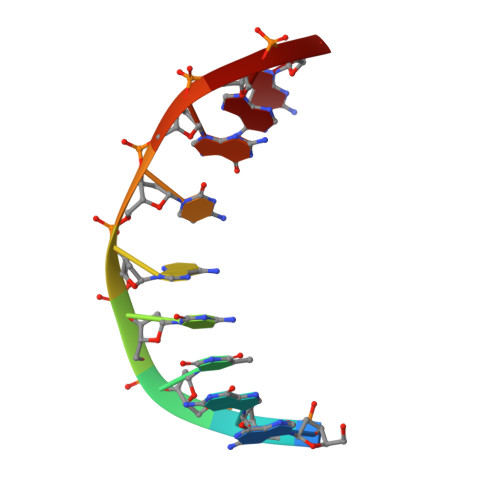
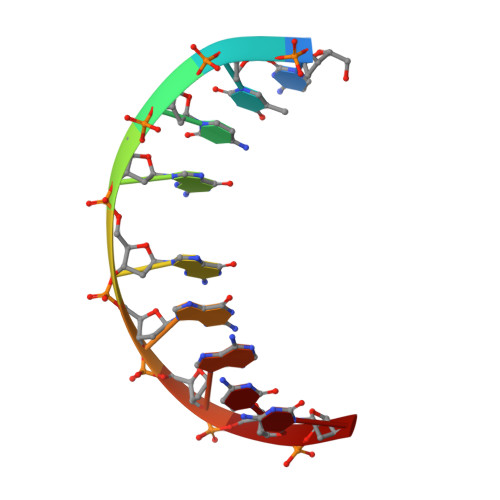
NMR solution structure of a nonanucleotide duplex with a dG mismatch opposite a 10S adduct derived from trans addition of a deoxyadenosine N6-amino group to (+)-(7R,8S,9S,10R)-7,8-dihydroxy-9,10-epoxy-7,8,9,10- tetrahydrobenzo[a]pyrene: an unusual syn glycosidic torsion angle at the modified dA
Yeh, H.J., Sayer, J.M., Liu, X., Altieri, A.S., Byrd, R.A., Lakshman, M.K., Yagi, H., Schurter, E.J., Gorenstein, D.G., Jerina, D.M.(1995) Biochemistry 34: 13570-13581
- PubMed: 7577946 Search on PubMed
- DOI: https://doi.org/10.1021/bi00041a037
- Primary Citation Related Structures:
1DXA - PubMed Abstract:
A nonanucleotide, d(G1G2T3C4[BaP]A5C6G7A8G9), in which (+)-(7R,8S,9S,10R)-7,8-dihydroxy-9,10-epoxy-7,8,9,10- tetrahydrobenzo[a]pyrene (7-hydroxyl group and epoxide oxygen are trans) is covalently bonded to the exocyclic N6-amino group of deoxyadenosine (dA5) through trans addition at C10 of the epoxide (to give a 10S adduct) has been synthesized. The solution structure of the duplex, d(G1G2T3C4[BaP]A5C6G7A8G9).d(C10T11C12G13G14G15A16C17C18+ ++), containing a dG mismatch opposite the modified dA (designated 10S-[BaP]dA.dG 9-mer duplex) has been investigated using a combination of 1D and 2D (including COSY, PECOSY, TOCSY, NOESY, and indirect detection of 1H-31P HETCOR) NMR spectroscopies. The NMR results together with restrained molecular dynamics/energy minimization calculations show that the modified dA5 adopts a syn glycosidic torsion angle whereas all other nucleotide residues adopt anti glycosidic torsion angles. The sugar ring of dA5 is in the C3'-endo conformation, and the sugar rings of the other residues are in the C2'-endo conformation. The hydrocarbon attached at dA5 orients toward the 3' end of the modified strand (i.e., dC6 direction) and intercalates between and parallel to bases of dG13 and dG14 of the complementary strand directly opposite dC6 and dA5, respectively. The edge of the hydrocarbon bearing H11 and H12 is positioned between the imino protons of dG13 and dG14 in the interior of the duplex, whereas H4 and H5 at the opposite edge are positioned near the sugar H1' and H2" protons of dG13 and facing the exterior of the duplex. The mismatched AG base pair is stabilized by dAsyn-dGanti base pairing in which the imino proton and the O6 of dG14 are hydrogen bonded to N7- and the single N6-amino proton, respectively, of the modified dA5. The modified DNA duplex remains in a right-handed helix, which bends at the site of intercalation about 20 to 30 degrees away from the helical axis and toward the direction of the modified strand.
- NIDDK, National Institutes of Health, Bethesda, Maryland 20892, USA.
Organizational Affiliation: