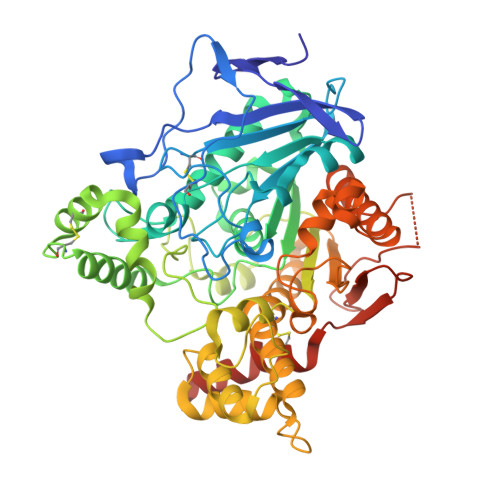
Structure of Acetylcholinesterase Complexed with (-)-Galanthamine at 2.3A Resolution
Greenblatt, H.M., Kryger, G., Lewis, T.T., Silman, I., Sussman, J.L.(1999) FEBS Lett 463: 321
- PubMed: 10606746 Search on PubMed
- DOI: https://doi.org/10.1016/s0014-5793(99)01637-3
- Primary Citation Related Structures:
1DX6 - PubMed Abstract:
(-)-Galanthamine (GAL), an alkaloid from the flower, the common snowdrop (Galanthus nivalis), shows anticholinesterase activity. This property has made GAL the target of research as to its effectiveness in the treatment of Alzheimer's disease. We have solved the X-ray crystal structure of GAL bound in the active site of Torpedo californica acetylcholinesterase (TcAChE) to 2.3 A resolution. The inhibitor binds at the base of the active site gorge of TcAChE, interacting with both the choline-binding site (Trp-84) and the acyl-binding pocket (Phe-288, Phe-290). The tertiary amine group of GAL does not interact closely with Trp-84; rather, the double bond of its cyclohexene ring stacks against the indole ring. The tertiary amine appears to make a non-conventional hydrogen bond, via its N-methyl group, to Asp-72, near the top of the gorge. The hydroxyl group of the inhibitor makes a strong hydrogen bond (2.7 A) with Glu-199. The relatively tight binding of GAL to TcAChE appears to arise from a number of moderate to weak interactions with the protein, coupled to a low entropy cost for binding due to the rigid nature of the inhibitor.
- Department of Structural Biology, Weizmann Institute of Science, Rehovot, Israel.
Organizational Affiliation: