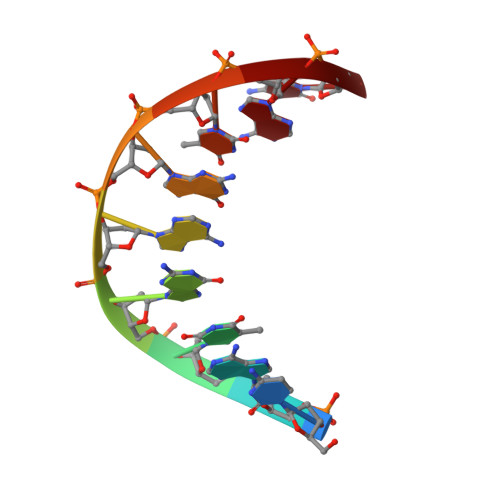
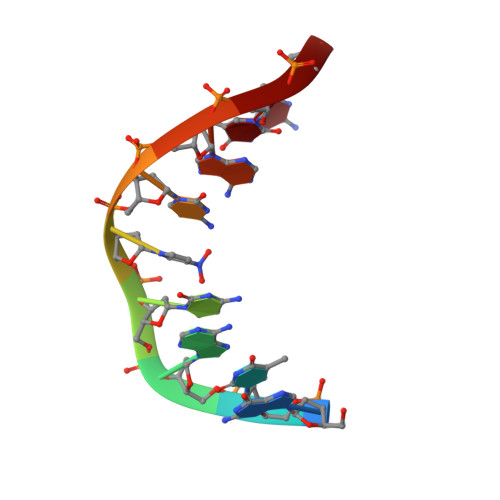
NMR structure of a DNA duplex containing nucleoside analog 1-(2'-deoxy-beta-D-ribofuranosyl)-3-nitropyrrole and the structure of the unmodified control.
Klewer, D.A., Hoskins, A., Zhang, P., Davisson, V.J., Bergstrom, D.E., LiWang, A.C.(2000) Nucleic Acids Res 28: 4514-4522
- PubMed: 11071940 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/28.22.4514
- Primary Citation Related Structures:
1DK6, 1DK9 - PubMed Abstract:
The three-dimensional structures of two DNA duplexes d(CATGAGTAC). d(GTACXCATG) (1) and d(CATGAGTAC).d(GTACTCATG) (2), where X represents 1-(2'-deoxy-beta-D-ribofuranosyl)-3-nitropyrrole, were solved using high resolution nuclear magnetic resonance spectroscopy and restrained molecular dynamics. Good convergence was observed between final structures derived from A- and B-form starting geometries for both 1 and 2. Structures of 1 and 2 are right-handed duplexes within the B-form conformational regime. Furthermore, the structures of 1 and 2 are highly similar, with differences in the structures localized to the vicinity of residue 14 (X versus T). The pyrrole group of 1 is in the syn conformation and it is displaced towards the major groove. Furthermore, unlike T14 in 2, the base of X14 has reduced pi-pi stacking interactions with C13 and C15 and the nitro group of X14 is pointing out of the major groove. The structures presented here establish the basis of the thermal data of DNA duplexes containing X and should be informative during the design of improved wild card nucleobase analogs.
- Department of Biochemistry and Biophysics, Texas A&M University, College Station, TX 77843-2128, USA.
Organizational Affiliation: