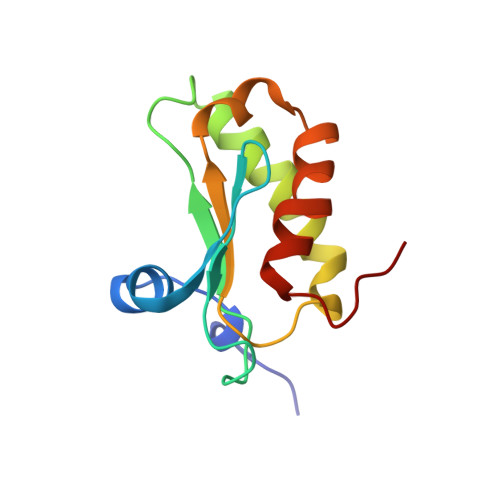
The structure of ribonuclease P protein from Staphylococcus aureus reveals a unique binding site for single-stranded RNA.
Spitzfaden, C., Nicholson, N., Jones, J.J., Guth, S., Lehr, R., Prescott, C.D., Hegg, L.A., Eggleston, D.S.(2000) J Mol Biology 295: 105-115
- PubMed: 10623511
- DOI: https://doi.org/10.1006/jmbi.1999.3341
- Primary Citation of Related Structures:
1D6T - PubMed Abstract:
Ribonuclease P (RNaseP) catalyses the removal of the 5'-leader sequence from pre-tRNA to produce the mature 5' terminus. The prokaryotic RNaseP holoenzyme consists of a catalytic RNA component and a protein subunit (RNaseP protein), which plays an auxiliary but essential role in vivo by binding to the 5'-leader sequence and broadening the substrate specificity of the ribozyme. We determined the three-dimensional high-resolution structure of the RNaseP protein from Staphylococcus aureus (117 amino acid residues) by nuclear magnetic resonance (NMR) spectroscopy in solution. The protein has an alphabeta-fold, similar to the ribonucleoprotein domain. We used small nucleic acid molecules as a model for the 5'-leader sequence to probe the propensity for generic single-stranded RNA binding on the protein surface. The NMR results reveal a contiguous interaction site, which is identical with the previously identified leader sequence binding site in RNaseP holoenzyme. The conserved arginine-rich motif does not bind single-stranded RNA. It is likely that this peptide segment binds selectively to double-stranded sections of P RNA, which are conformationally more rigid. Given the essentiality of RNaseP for the viability of the organism, knowledge of the S. aureus protein structure and insight into its interaction with RNA will help us to develop RNaseP and RNaseP protein as targets for novel antibiotics against this pathogen.
- Computational and Structural Sciences, SmithKline Beecham Pharmaceuticals, Harlow, CM19 5AW, UK. Claus_Spitzfaden-1@sbphrd.com
Organizational Affiliation: