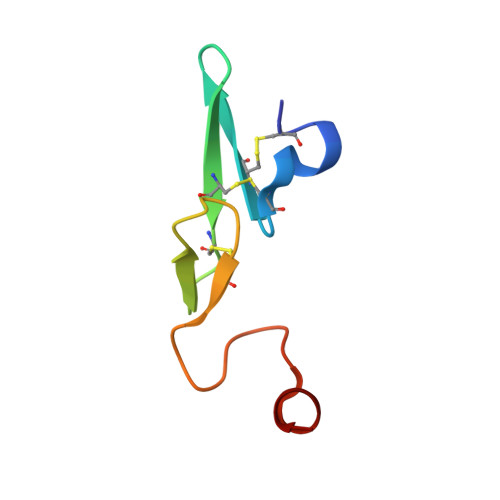
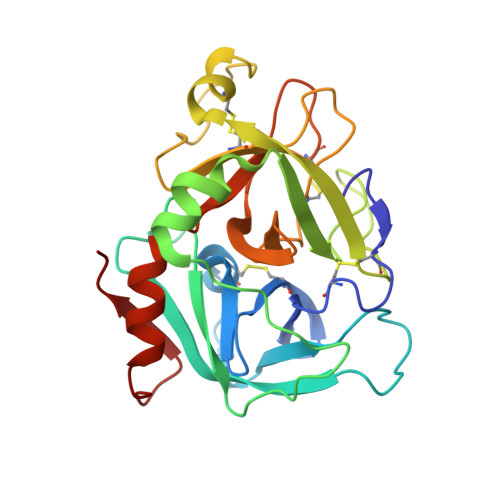
Crystal structure of active site-inhibited human coagulation factor VIIa (des-Gla).
Kemball-Cook, G., Johnson, D.J., Tuddenham, E.G., Harlos, K.(1999) J Struct Biol 127: 213-223
- PubMed: 10544046 Search on PubMed
- DOI: https://doi.org/10.1006/jsbi.1999.4158
- Primary Citation Related Structures:
1CVW - PubMed Abstract:
Factor VIIa (FVIIa) is a crucial haemostatic protease consisting of four distinct domains termed the Gla, epidermal growth factor-1 (EGF-1), EGF-2, and protease domains (from N- to C-terminus). The crystal structure of human FVIIa inhibited at the active site with 1, 5-dansyl-Glu-Gly-Arg-chloromethyl ketone and lacking the Gla domain has been solved to a resolution of 2.28 A. The EGF-2 and protease domains were well resolved, whereas no electron density for the EGF-1 domain was observed, suggesting a flexible arrangement or disorder within the crystal. Superposition of the protease domain of the present structure with that previously resolved in the tissue factor (TF)/FVIIai complex revealed that although overall the domain structures are similar, the EGF-2 domain is rotated by 7.5 degrees relative to the protease domain on binding TF. A single cleavage in the protease domain was found, between Arg315 and Lys316 (chymotrypsin numbering 170C-170D) in a FVII-specific insertion loop: this cleavage appeared to be essential for crystallisation. Insertion of the heavy chain N-terminal Ile153 is essentially identical in the two structures, as is the geometry of the active site residues and the inhibitor C-terminal arginine residue. Some differences are seen in the cleaved loop, but changes in TF-contact residues are generally minor. This structure supports the hypothesis that TF binding enables spatial domain arrangements in the flexible FVIIa molecule necessary for procoagulant function and furthermore that active site occupancy induces FVIIa active conformation via N-terminal insertion.
- MRC Clinical Sciences Centre, Imperial College School of Medicine, Hammersmith Hospital, Du Cane Road, London, W12 0NN, United Kingdom. gkemball@hgmp.mrc.ac.uk
Organizational Affiliation: