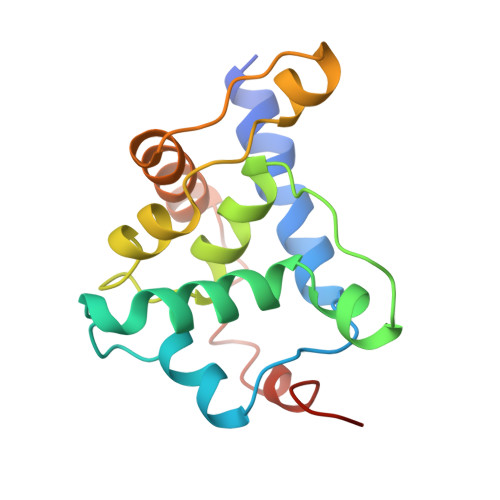
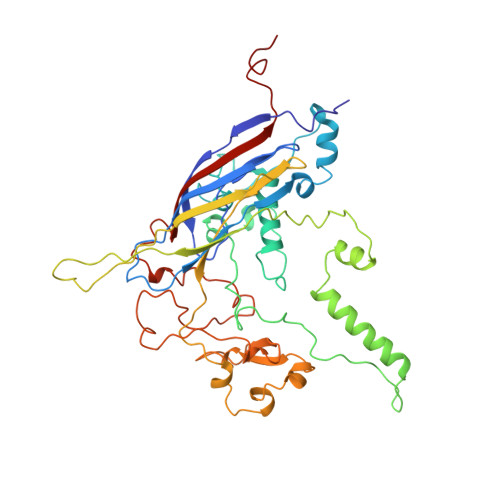
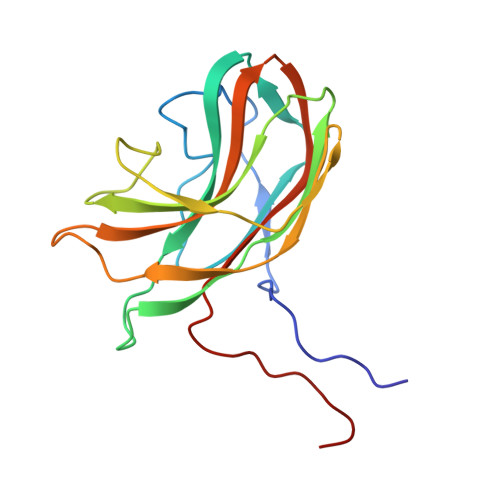
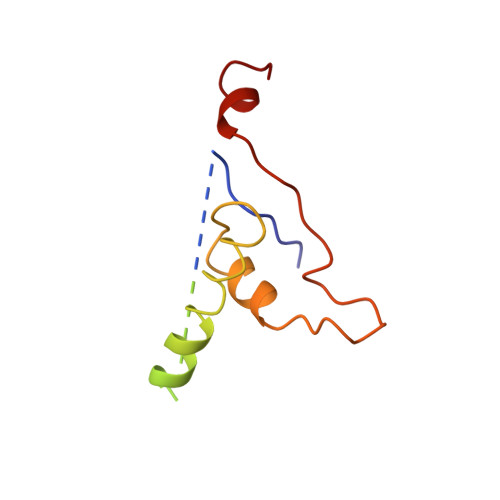
The role of scaffolding proteins in the assembly of the small, single-stranded DNA virus phiX174.
Dokland, T., Bernal, R.A., Burch, A., Pletnev, S., Fane, B.A., Rossmann, M.G.(1999) J Mol Biology 288: 595-608
- PubMed: 10329166 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1999.2699
- Primary Citation Related Structures:
1CD3 - PubMed Abstract:
An empty precursor particle called the procapsid is formed during assembly of the single-stranded DNA bacteriophage phiX174. Assembly of the phiX174 procapsid requires the presence of the two scaffolding proteins, D and B, which are structural components of the procapsid, but are not found in the mature virion. The X-ray crystallographic structure of a "closed" procapsid particle has been determined to 3.5 A resolution. This structure has an external scaffold made from 240 copies of protein D, 60 copies of the internally located B protein, and contains 60 copies of each of the viral structural proteins F and G, which comprise the shell and the 5-fold spikes, respectively. The F capsid protein has a similar conformation to that seen in the mature virion, and differs from the previously determined 25 A resolution electron microscopic reconstruction of the "open" procapsid, in which the F protein has a different conformation. The D scaffolding protein has a predominantly alpha-helical fold and displays remarkable conformational variability. We report here an improved and refined structure of the closed procapsid and describe in some detail the differences between the four independent D scaffolding proteins per icosahedral asymmetric unit, as well as their interaction with the F capsid protein. We re-analyze and correct the comparison of the closed procapsid with the previously determined cryo-electron microscopic image reconstruction of the open procapsid and discuss the major structural rearrangements that must occur during assembly. A model is proposed in which the D proteins direct the assembly process by sequential binding and conformational switching.
- Department of Biological Sciences, Purdue University, West Lafayette, IN, 47907-1392, USA.
Organizational Affiliation: