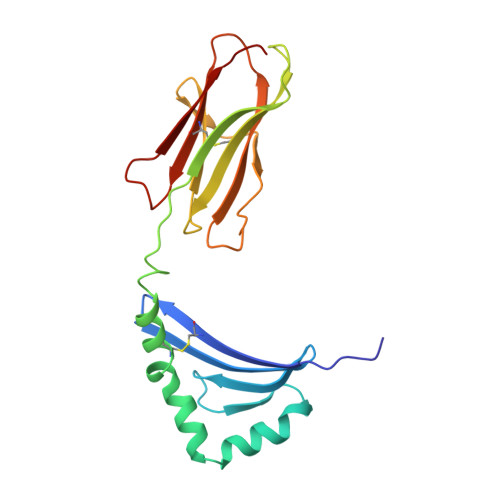
Crystal structure of HLA-DR2 (DRA*0101, DRB1*1501) complexed with a peptide from human myelin basic protein.
Smith, K.J., Pyrdol, J., Gauthier, L., Wiley, D.C., Wucherpfennig, K.W.(1998) J Exp Medicine 188: 1511-1520
- PubMed: 9782128
- DOI: https://doi.org/10.1084/jem.188.8.1511
- Primary Citation of Related Structures:
1BX2 - PubMed Abstract:
Susceptibility to multiple sclerosis is associated with the human histocompatibility leukocyte antigen (HLA)-DR2 (DRB1*1501) haplotype. The structure of HLA-DR2 was determined with a bound peptide from human myelin basic protein (MBP) that is immunodominant for human MBP-specific T cells. Residues of MBP peptide that are important for T cell receptor recognition are prominent, solvent exposed residues in the crystal structure. A distinguishing feature of the HLA-DR2 peptide binding site is a large, primarily hydrophobic P4 pocket that accommodates a phenylalanine of the MBP peptide. The necessary space for this aromatic side chain is created by an alanine at the polymorphic DRbeta 71 position. These features make the P4 pocket of HLA-DR2 distinct from DR molecules associated with other autoimmune diseases.
- Department of Molecular Medicine, Children's Hospital, Boston, Massachusetts 02115, USA. ksmith@rascal.med.harvard.edu
Organizational Affiliation: