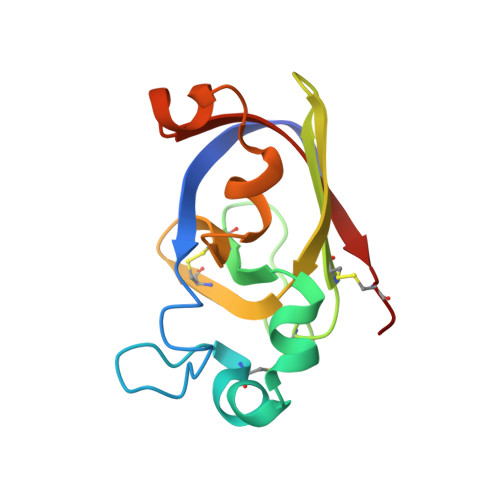
Three-dimensional structure in solution of barwin, a protein from barley seed.
Ludvigsen, S., Poulsen, F.M.(1992) Biochemistry 31: 8783-8789
- PubMed: 1390665 Search on PubMed
- DOI: https://doi.org/10.1021/bi00152a014
- Primary Citation Related Structures:
1BW3, 1BW4 - PubMed Abstract:
The solution structure of a 125-residue basic protein, barwin, has been determined using 1H nuclear magnetic resonance spectroscopy. This protein is closely related to domains in proteins encoded by wound-induced genes in plants. Analysis of the 1H nuclear Overhauser spectrum revealed the assignment of more than 1400 nuclear Overhauser effects. Twenty structures were calculated based on 676 nontrivial distance restraints, 152 torsion angle restraints (92 phi, 56 chi 1, and 4 omega for proline), and stereospecific assignments of 38 chiral centers, using distance geometry, simulated annealing, and restrained energy minimization. None of the distance restraints was violated by more than 0.5 A in any of the 20 structures, and none of the torsion angle restraints was violated by more than 1 degree in any of the structures. The RMS difference between the calculated and target interproton distance restraints is 0.033 A, and the average atomic RMS differences between the 20 structures and their geometric average are 1.23 A for backbone atoms and 1.73 A for all heavy atoms. The dominating structural feature of the protein is a well-defined four-stranded antiparallel beta-sheet, two parallel beta-sheets packed antiparallel to each other and four short alpha-helices. The binding site of barwin to the tetramer N-acetylglucosamine has been qualitatively investigated, and the dissociation constant of the complex has been determined using one-dimensional 1H nuclear magnetic resonance spectroscopy.
- Carlsberg Laboratorium, Kemisk Afdeling, Copenhagen, Denmark.
Organizational Affiliation: