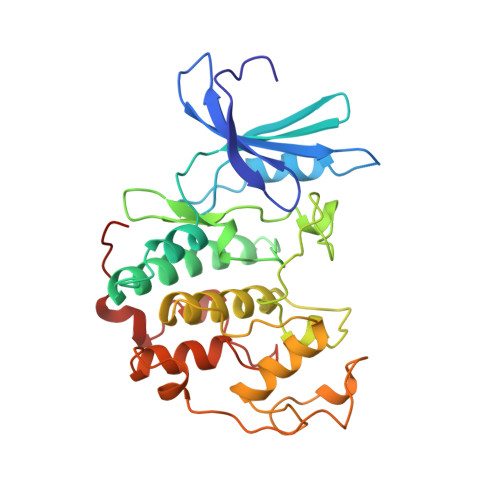
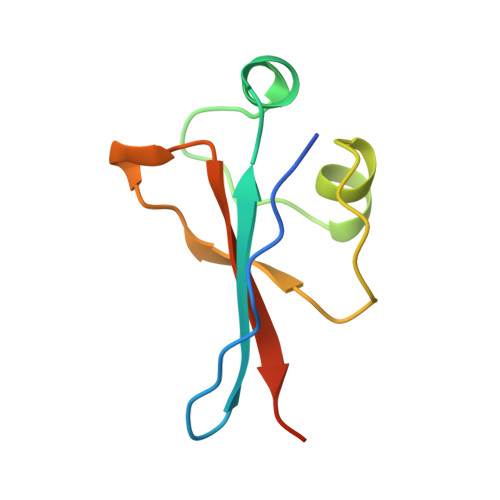
Crystal structure and mutational analysis of the human CDK2 kinase complex with cell cycle-regulatory protein CksHs1.
Bourne, Y., Watson, M.H., Hickey, M.J., Holmes, W., Rocque, W., Reed, S.I., Tainer, J.A.(1996) Cell 84: 863-874
- PubMed: 8601310 Search on PubMed
- DOI: https://doi.org/10.1016/s0092-8674(00)81065-x
- Primary Citation Related Structures:
1BUH - PubMed Abstract:
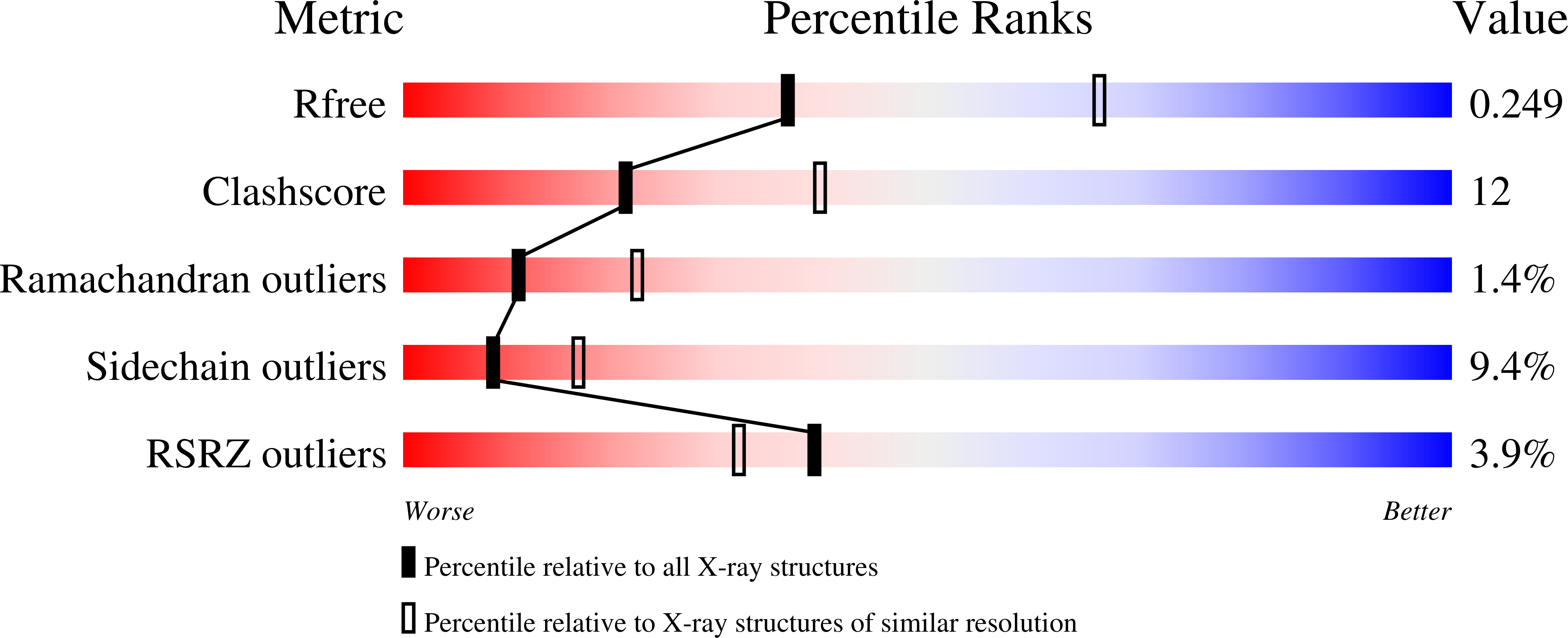
The 2.6 Angstrom crystal structure for human cyclin-dependent kinase 2(CDK2) in complex with CksHs1, a human homolog of essential yeast cell cycle-regulatory proteins suc1 and Cks1, reveals that CksHs1 binds via all four beta strands to the kinase C-terminal lobe. This interface is biologically critical, based upon mutational analysis, but far from the CDK2 N-terminal lobe, cyclin, and regulatory phosphorylation sites. CDK2 binds the Cks single domain conformation and interacts with conserved hydrophobic residues plus His-60 and Glu-63 in their closed beta-hinge motif conformation. The beta hinge opening to form the Cks beta-interchanged dimer sterically precludes CDK2 binding, providing a possible mechanism regulating CDK2-Cks interactions. One face of the complex exposes the sequence-conserved phosphate-binding region on Cks and the ATP-binding site on CDK2, suggesting that CKs may target CDK2 to other phosphoproteins during the cell cycle.
- Department of Molecular Biology, Scripps Research Institute, La Jolla, California, 92037, USA.
Organizational Affiliation: