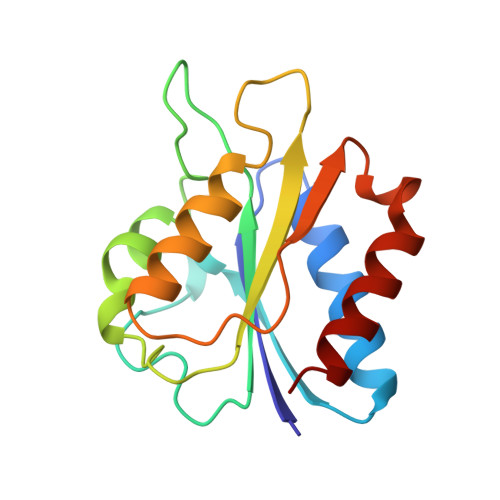
X-ray crystal structure of the Desulfovibrio vulgaris (Hildenborough) apoflavodoxin-riboflavin complex.
Walsh, M.A., McCarthy, A., O'Farrell, P.A., McArdle, P., Cunningham, P.D., Mayhew, S.G., Higgins, T.M.(1998) Eur J Biochem 258: 362-371
- PubMed: 9874201 Search on PubMed
- DOI: https://doi.org/10.1046/j.1432-1327.1998.2580362.x
- Primary Citation Related Structures:
1BU5 - PubMed Abstract:
The apoprotein of flavodoxin from Desulfovibrio vulgaris forms a complex with riboflavin. The ability to bind riboflavin distinguishes this flavodoxin from other short-chain flavodoxins which require the phosphate of FMN for flavin binding. The redox potential of the semiquinone/hydroquinone couple of the bound riboflavin is 180 mV less negative than the corresponding complex with FMN. To elucidate the binding of riboflavin, the complex has been crystallized and the crystal structure solved by molecular replacement using native flavodoxin as a search model to a resolution of 0.183 nm. Compared to the FMN complex, the hydrogen-bonding network at the isoalloxazine sub-site of the riboflavin complex is severely disrupted by movement of the loop residues Ser58-Ile64 (60-loop) which contact the isoalloxazine by over 0.35 nm, and by a small displacement of the isoalloxazine moiety. The 60-loop movement away from the flavin increases the solvent exposure of the flavin-binding site. The conformation of the site at which 5'-phosphate of FMN normally binds is similar in the two complexes, but in the riboflavin complex a sulphate or phosphate ion from the crystallization buffer occupies the space. This causes small structural perturbations in the phosphate-binding site. The flexibility of the 60-loop in D. vulgaris flavodoxin appears to be a contributing factor to the binding of riboflavin by the apoprotein, and a feature that distinguishes the protein from other 'short chain' flavodoxins. In the absence of the terminal phosphate group, free movement at the 5'-OH group of the ribityl chain can occur. Thus, the 5'-phosphate of FMN secures the cofactor at the binding site and positions it optimally. The structural changes which occur in the 60-loop in the riboflavin complex probably account for most of the positive shift that is observed in the midpoint potential of the semiquinone/hydroquinone couple of the riboflavin complex compared to that of the FMN complex.
- Department of Chemistry, University College, Galway, Ireland. walsh@anl.gov
Organizational Affiliation: