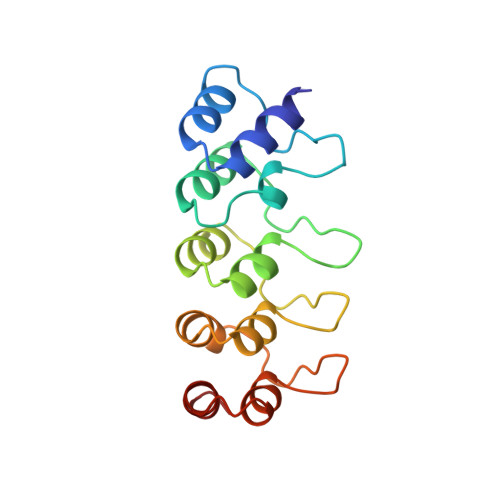
Structure of human cyclin-dependent kinase inhibitor p19INK4d: comparison to known ankyrin-repeat-containing structures and implications for the dysfunction of tumor suppressor p16INK4a.
Baumgartner, R., Fernandez-Catalan, C., Winoto, A., Huber, R., Engh, R.A., Holak, T.A.(1998) Structure 6: 1279-1290
- PubMed: 9782052 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(98)00128-2
- Primary Citation Related Structures:
1BD8 - PubMed Abstract:
The four members of the INK4 gene family (p16(INK4a), p15(INK4b), p18(INK4c) and p19(INK4d)) inhibit the closely related cyclin-dependent kinases CDK4 and CDK6 as part of the regulation of the G1-->S transition in the cell-division cycle. Loss of INK4 gene product function, particularly that of p16(INK4a), is found in 10-60% of human tumors, suggesting that broadly applicable anticancer therapies might be based on restoration of p16(INK4a) CDK inhibitory function. Although much less frequent, defects of p19(INK4d) have also been associated with human cancer (osteosarcomas). The protein structures of some INK4 family members, determined by nuclear magnetic resonance (NMR) spectroscopy and X-ray techniques, have begun to clarify the functional role of p16(INK4a) and the dysfunction introduced by the mutations associated with human tumors. The crystal structure of human p19(INK4d) has been determined at 1.8 A resolution using multiple isomorphous replacement methods. The fold of p19(INK4d) produces an oblong molecule comprising five approximately 32-residue ankyrin-like repeats. The architecture of the protein demonstrates the high structural similarity within the INK4 family. Comparisons to other ankyrin-repeat-containing proteins (GABPbeta, 53BP2 and myotrophin) show similar structures with comparable hydrogen-bonding patterns and hydrophobic interactions. Such comparisons highlight the splayed beta-loop geometry that is specific to INK4 inhibitors. This geometry is the result of a modified ankyrin structure in the second repeat. Among the INK4 inhibitors, the highest amino acid sequence conservation is found in the helical stacks; this conservation creates a conserved beta-loop geometry specific to INK4 inhibitors. Therefore, in addition to models which predict that the conserved helix alpha6 is responsible for CDK inhibition, a binding mode whereby the loops of INK4 proteins bind to the CDKs should also be considered. A similar loop-based interaction is seen in the complex formed between the ankyrin-repeat-containing protein GABPbeta and_GABPalpha. This mode of binding would be consistent with the observation that p16(INK4a) is sensitive to deleterious mutations found throughout this tumor suppressor protein; these mutations probably destabilize the three-dimensional structure.
- Max Planck Institute for Biochemistry D-82152, Martinsried, Federal Republic of Germany.
Organizational Affiliation: