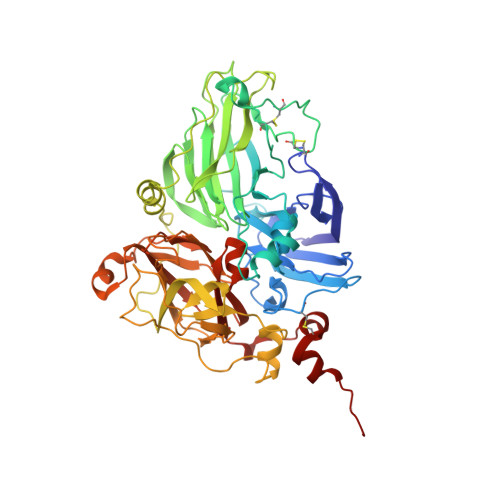
X-ray structures and mechanistic implications of three functional derivatives of ascorbate oxidase from zucchini. Reduced, peroxide and azide forms.
Messerschmidt, A., Luecke, H., Huber, R.(1993) J Mol Biology 230: 997-1014
- PubMed: 8478945
- DOI: https://doi.org/10.1006/jmbi.1993.1215
- Primary Citation of Related Structures:
1ASO, 1ASP, 1ASQ - PubMed Abstract:
The X-ray structures of three functional derivatives of ascorbate oxidase (EC 1.10.3.3) from Zucchini have been determined and are compared to the "native" oxidized form. The fully reduced form of ascorbate oxidase has been refined to a crystallographic R-factor of 19.6% for all reflections between 8.0 A and 2.2 A resolution. The geometry at the type-1 copper (CU1) is unchanged compared to the oxidized form, but the oxygen ligand bridging the copper ions CU2 and CU3 (spectroscopic type-3 copper pair) is released and the copper ions move apart yielding a trigonal planar co-ordination with their ligating histidine residues. The co-ordination at the copper ion CU4 (spectroscopic type-2 copper) is not affected. The copper-copper distances increase from an average 3.7 A in the native form to 5.1 A for CU2-CU3, 4.4 A for CU2-CU4 and 4.1 A for CU3-CU4. The peroxide derivative of ascorbate oxidase has been refined to a crystallographic R-factor of 16.0% for all reflections between 8.0 A and 2.59 A resolution. The geometry at the type-1 copper site is not changed compared to the oxidized form. The oxygen ligand bridging copper atoms CU2 and CU3 is lost, too. The peroxide binds terminally to the copper ion CU2 as hydroperoxide. Copper ion CU2 is fourfold co-ordinated to the NE2 atoms of the three histidine residues and to the oxygen atom of the terminally bound peroxide molecule in a distorted tetrahedral geometry. Copper ion CU3 is threefold co-ordinated as in the reduced form and co-ordination around copper atom CU4 is unaltered. The copper-copper distances increase to 4.8 A for CU2-CU3 and 4.5 A for CU2-CU4. The distance CU3-CU4 remains 3.7 A. Treatment with peroxide causes a partial depletion of copper ion CU2. The refinement for the azide derivative of ascorbate oxidase converged at a crystallographic R-factor of 17.8% for all reflections between 8.0 A and 2.32 A. There are no significant structural changes at the type-1 copper site. The oxygen ligand bridging copper ions CU2 and CU3 is again released. Two azide molecules bind terminally to copper ion CU2. Copper ion CU2 is fivefold co-ordinated to the NE2 atoms of the three histidine residues and to both terminally bound azide molecules in a trigonal-bipyramidal manner. Copper-copper distances increase to 5.1 A for CU2-CU3 and 4.6 A for CU2-CU4. The distance CU3-CU4 is decreased to 3.6 A.(ABSTRACT TRUNCATED AT 400 WORDS)
- Max-Planck-Institut für Biochemie, Martinsried, FRG.
Organizational Affiliation: