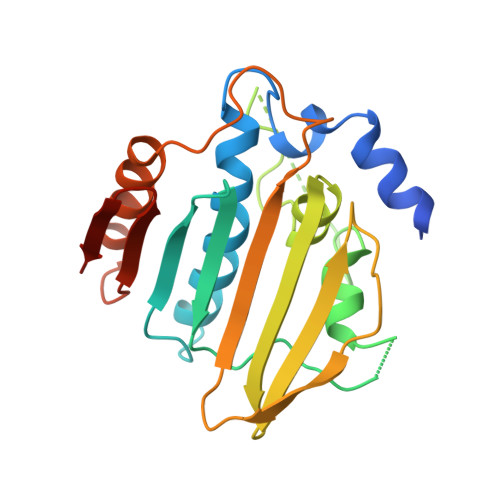
The entropic penalty of ordered water accounts for weaker binding of the antibiotic novobiocin to a resistant mutant of DNA gyrase: a thermodynamic and crystallographic study.
Holdgate, G.A., Tunnicliffe, A., Ward, W.H., Weston, S.A., Rosenbrock, G., Barth, P.T., Taylor, I.W., Pauptit, R.A., Timms, D.(1997) Biochemistry 36: 9663-9673
- PubMed: 9245398
- DOI: https://doi.org/10.1021/bi970294+
- Primary Citation of Related Structures:
1AJ6 - PubMed Abstract:
Novobiocin is an antibiotic which binds to a 24 kDa fragment from the B subunit of DNA gyrase. Naturally occurring resistance arises from mutation of Arg-136 which hydrogen bonds to the coumarin ring of novobiocin. We have applied calorimetry to characterize the binding of novobiocin to wild-type and R136H mutant 24 kDa fragments. Upon mutation, the Kd increases from 32 to 1200 nM at 300 K. The enthalpy of binding is more favorable for the mutant (DeltaH degrees shifts from -12.1 to -17.5 kcal/mol), and the entropy of binding is much less favorable (TDeltaS degrees changes from -1.8 to -9.4 kcal/mol). Both of these changes are in the direction opposite to that expected if the loss of the Arg residue reduces hydrogen bonding. The change in heat capacity at constant pressure upon binding (DeltaCp) shifts from -295 to -454 cal mol-1 K-1. We also report the crystal structure, at 2.3 A resolution, of a complex between the R136H 24 kDa fragment and novobiocin. Although the change in DeltaCp often would be interpreted as reflecting increased burial of hydrophobic surface on binding, this structure reveals a small decrease. Furthermore, an ordered water molecule is sequestered into the volume vacated by removal of the guanidinium group. There are large discrepancies when the measured thermodynamic parameters are compared to those estimated from the structural data using empirical relationships. These differences seem to arise from the effects of sequestering ordered water molecules upon complexation. The water-mediated hydrogen bonds linking novobiocin to the mutant protein make a favorable enthalpic contribution, whereas the immobilization of the water leads to an entropic cost and a reduction in the heat capacity of the system. Such a negative contribution to DeltaCp, DeltaH degrees , and TDeltaS degrees appears to be a general property of water molecules that are sequestered when ligands bind to proteins.
- ZENECA Pharmaceuticals, Macclesfield, Cheshire, U.K.
Organizational Affiliation: