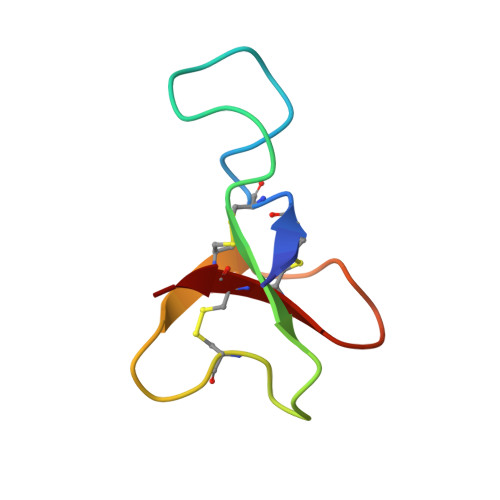
Three-dimensional structure in solution of the polypeptide cardiac stimulant anthopleurin-A.
Pallaghy, P.K., Scanlon, M.J., Monks, S.A., Norton, R.S.(1995) Biochemistry 34: 3782-3794
- PubMed: 7893675
- DOI: https://doi.org/10.1021/bi00011a036
- Primary Citation Related Structures:
1AHL - PubMed Abstract:
The three-dimensional structure in aqueous solution of the 49-residue polypeptide anthopleurin-A (AP-A), from the sea anemone Anthopleura xanthogrammica, has been determined from 1H NMR data. A restraint set consisting of 411 interproton distance restraints inferred from NOEs and 19 backbone and 13 side chain dihedral angle restraints from spin-spin coupling constants, as well as 15 lower bound restraints based on the absence of NOEs in the spectra, was used as input for distance geometry calculations in DIANA and simulated annealing and restrained energy minimization in X-PLOR. Stereospecific assignments for 12 beta-methylene pairs were also included. The final set of 20 structures had mean pairwise rms differences over the whole molecule of 2.04 A for the backbone heavy atoms (N, C alpha, and C) and 2.59 A for all heavy atoms. For the well-defined region encompassing residues 2-7 and 17-49, the corresponding values were 0.82 and 1.27 A, respectively. AP-A adopts a compact structure consisting of four short strands of antiparallel beta-sheet (residues 2-4, 20-23, 34-37, and 45-48) connected by three loops. The first loop commences with a type I beta-turn which includes two important Asp residues; this loop is the least well-defined region of the protein, although a beta-turn involving residues 13-16 is observed in nearly half the structures. The loop linking the second and third strands is constrained by the 29-47 disulfide bond and contains two well-defined beta-turns, while the third loop contains the Gly40-Pro41 sequence, which has been identified previously as the site of cis-trans isomerism. The carboxylate group of Asp7 is close to the epsilon-ammonium group of Lys37, suggesting that they may form a salt bridge. A pH titration monitored by 2D NMR supports this by showing that Asp7 has a low pKa. It is proposed that this region of the molecule and the nearby residues Asp9 and His39 form part of the molecular surface which interacts with the mammalian cardiac sodium channel.
- NMR Laboratory, Biomolecular Research Institute, Parkville, Australia.
Organizational Affiliation: