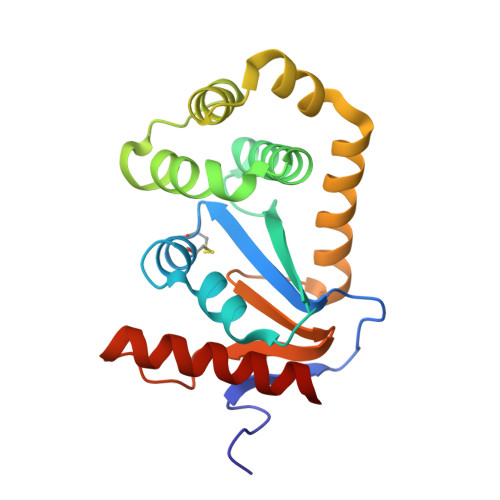
Structural analysis of three His32 mutants of DsbA: support for an electrostatic role of His32 in DsbA stability.
Guddat, L.W., Bardwell, J.C., Glockshuber, R., Huber-Wunderlich, M., Zander, T., Martin, J.L.(1997) Protein Sci 6: 1893-1900
- PubMed: 9300489 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.5560060910
- Primary Citation Related Structures:
1AC1, 1ACV, 1FVJ, 1FVK - PubMed Abstract:
DsbA, a 21-kDa protein from Escherichia coli, is a potent oxidizing disulfide catalyst required for disulfide bond formation in secreted proteins. The active site of DsbA is similar to that of mammalian protein disulfide isomerases, and includes a reversible disulfide bond formed from cysteines separated by two residues (Cys30-Pro31-His32-Cys33). Unlike most protein disulfides, the active-site disulfide of DsbA is highly reactive and the oxidized form of DsbA is much less stable than the reduced form at physiological pH. His32, one of the two residues between the active-site cysteines, is critical to the oxidizing power of DsbA and to the relative instability of the protein in the oxidized form. Mutation of this single residue to tyrosine, serine, or leucine results in a significant increase in stability (of approximately 5-7 kcal/mol) of the oxidized His32 variants relative to the oxidized wild-type protein. Despite the dramatic changes in stability, the structures of all three oxidized DsbA His32 variants are very similar to the wild-type oxidized structure, including conservation of solvent atoms near the active-site residue, Cys30. These results show that the His32 residue does not exert a conformational effect on the structure of DsbA. The destabilizing effect of His32 on oxidized DsbA is therefore most likely electrostatic in nature.
- Centre for Drug Design and Development, University of Queensland, Brisbane, Australia.
Organizational Affiliation: