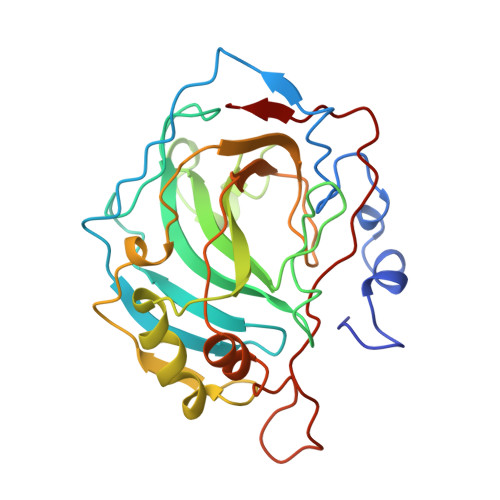
Altering the mouth of a hydrophobic pocket. Structure and kinetics of human carbonic anhydrase II mutants at residue Val-121.
Nair, S.K., Calderone, T.L., Christianson, D.W., Fierke, C.A.(1991) J Biological Chem 266: 17320-17325
- PubMed: 1910042 Search on PubMed
- Primary Citation Related Structures:
12CA - PubMed Abstract:
Eleven amino acid substitutions at Val-121 of human carbonic anhydrase II including Gly, Ala, Ser, Leu, Ile, Lys, and Arg, were constructed by site-directed mutagenesis. This residue is at the mouth of the hydrophobic pocket in the enzyme active site. The CO2 hydrase activity and the p-nitrophenyl esterase activity of these CAII variants correlate with the hydrophobicity of the residue, suggesting that the hydrophobic character of this residue is important for catalysis. The effects of these mutations on the steady-state kinetics for CO2 hydration occur mainly in kcat/Km and Km, consistent with involvement of this residue in CO2 association. The Val-121----Ala mutant, which exhibits about one-third normal CO2 hydrase activity, has been studied by x-ray crystallographic methods. No significant changes in the mutant enzyme conformation are evident relative to the wild-type enzyme. Since Val-121 is at the mouth of the hydrophobic pocket, its substitution by the methyl side chain of alanine makes the pocket mouth significantly wider than that of the wild-type enzyme. Hence, although a moderately wide (and deep) pocket is important for substrate association, a wider mouth to this pocket does not seriously compromise the catalytic approach of CO2 toward nucleophilic zinc-bound hydroxide.
- Department of Chemistry, University of Pennsylvania, Philadelphia 19104-6323.
Organizational Affiliation: