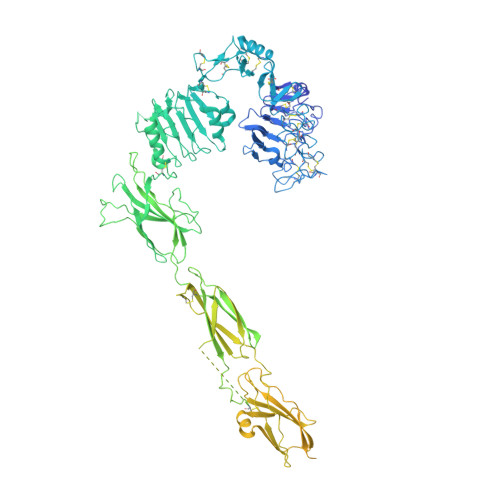
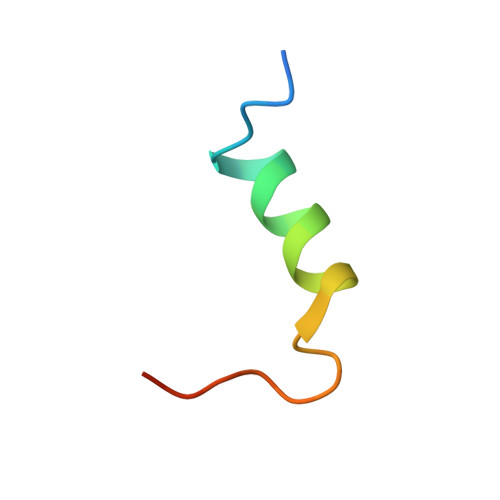
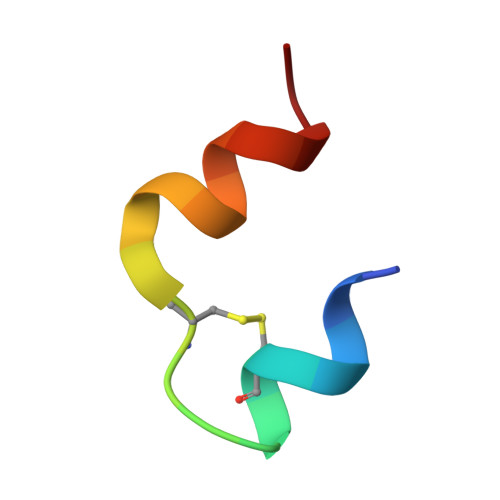Structural Investigations of Full-Length Insulin Receptor Dynamics and Signalling.
Nielsen, J., Brandt, J., Boesen, T., Hummelshoj, T., Slaaby, R., Schluckebier, G., Nissen, P.(2022) J Mol Biol 434: 167458-167458
- PubMed: 35074483
- DOI: https://doi.org/10.1016/j.jmb.2022.167458
- Primary Citation of Related Structures:
7PG0, 7PG2, 7PG3, 7PG4 - PubMed Abstract:
Insulin regulates glucose homeostasis via binding and activation of the insulin receptor dimer at two distinct pairs of binding sites 1 and 2. Here, we present cryo-EM studies of full-length human insulin receptor (hIR) in an active state obtained at non-saturating, physiologically relevant insulin conditions. Insulin binds asymmetrically to the receptor under these conditions, occupying up to three of the four possible binding sites. Deletion analysis of the receptor together with site specific peptides and insulin analogs used in binding studies show that both sites 1 and 2 are required for high insulin affinity. We identify a homotypic interaction of the fibronectin type III domain (FnIII-3) of IR resulting in tight interaction of membrane proximal domains of the active, asymmetric receptor dimer. Our results show how insulin binding at two distinct types of sites disrupts the autoinhibited apo-IR dimer and stabilizes the active dimer. We propose an insulin binding and activation mechanism, which is sequential, exhibits negative cooperativity, and is based on asymmetry at physiological insulin concentrations with one to three insulin molecules activating IR.
Organizational Affiliation:
Department of Molecular Biology and Genetics, Danish Research Institute of Translational Neuroscience - DANDRITE, Nordic EMBL Partnership for Molecular Medicine, Aarhus University, Aarhus, Denmark; Novo Nordisk A/S, Novo Nordisk Park, Måløv, Denmark.