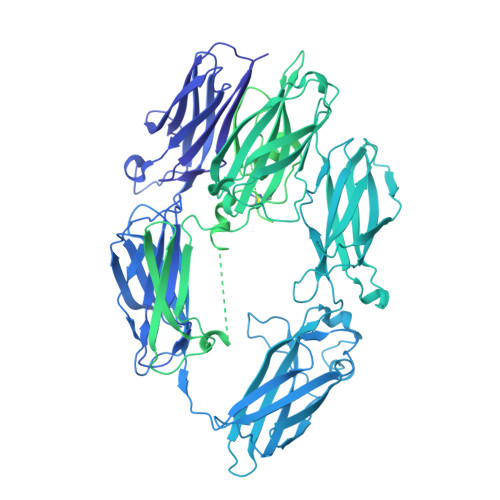
The allosteric modulation of Complement C5 by knob domain peptides.
Macpherson, A., Laabei, M., Ahdash, Z., Graewert, M.A., Birtley, J.R., Schulze, M.E., Crennell, S., Robinson, S.A., Holmes, B., Oleinikovas, V., Nilsson, P.H., Snowden, J., Ellis, V., Mollnes, T.E., Deane, C.M., Svergun, D., Lawson, A.D., van den Elsen, J.M.(2021) Elife 10
- PubMed: 33570492
- DOI: https://doi.org/10.7554/eLife.63586
- Primary Citation of Related Structures:
7AD6, 7AD7 - PubMed Abstract:
Bovines have evolved a subset of antibodies with ultra-long heavy chain complementarity determining regions that harbour cysteine-rich knob domains. To produce high-affinity peptides, we previously isolated autonomous 3-6 kDa knob domains from bovine antibodies. Here, we show that binding of four knob domain peptides elicits a range of effects on the clinically validated drug target complement C5. Allosteric mechanisms predominated, with one peptide selectively inhibiting C5 cleavage by the alternative pathway C5 convertase, revealing a targetable mechanistic difference between the classical and alternative pathway C5 convertases. Taking a hybrid biophysical approach, we present C5-knob domain co-crystal structures and, by solution methods, observed allosteric effects propagating >50 Å from the binding sites. This study expands the therapeutic scope of C5, presents new inhibitors, and introduces knob domains as new, low molecular weight antibody fragments, with therapeutic potential.
- UCB, Slough, United Kingdom.
Organizational Affiliation: