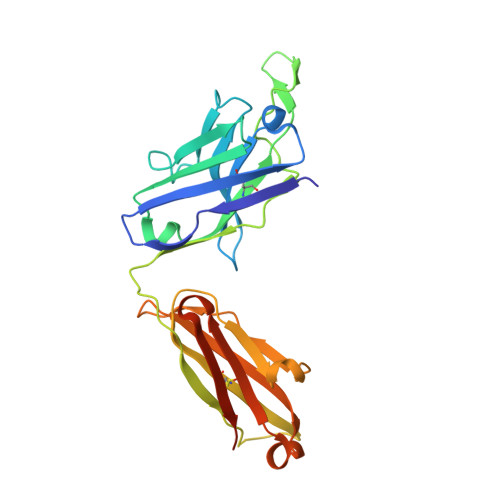
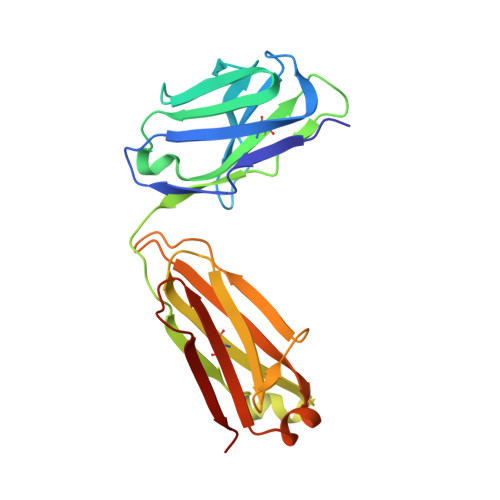
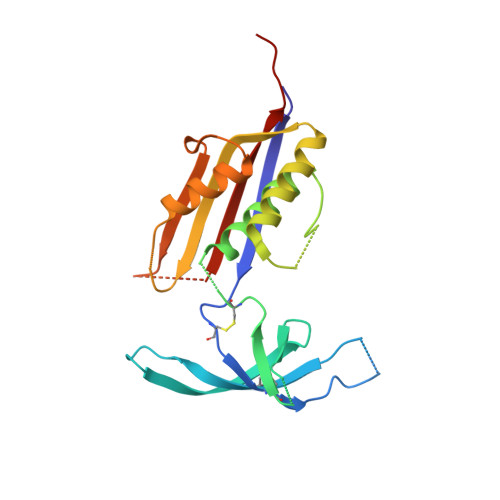
Structures of HIV-1 Env V1V2 with broadly neutralizing antibodies reveal commonalities that enable vaccine design.
Gorman, J., Soto, C., Yang, M.M., Davenport, T.M., Guttman, M., Bailer, R.T., Chambers, M., Chuang, G.Y., DeKosky, B.J., Doria-Rose, N.A., Druz, A., Ernandes, M.J., Georgiev, I.S., Jarosinski, M.C., Joyce, M.G., Lemmin, T.M., Leung, S., Louder, M.K., McDaniel, J.R., Narpala, S., Pancera, M., Stuckey, J., Wu, X., Yang, Y., Zhang, B., Zhou, T., Program, N.C., Mullikin, J.C., Baxa, U., Georgiou, G., McDermott, A.B., Bonsignori, M., Haynes, B.F., Moore, P.L., Morris, L., Lee, K.K., Shapiro, L., Mascola, J.R., Kwong, P.D.(2016) Nat Struct Mol Biol 23: 81-90
- PubMed: 26689967
- DOI: https://doi.org/10.1038/nsmb.3144
- Primary Citation of Related Structures:
5ESV, 5ESZ - PubMed Abstract:
Broadly neutralizing antibodies (bNAbs) against HIV-1 Env V1V2 arise in multiple donors. However, atomic-level interactions had previously been determined only with antibodies from a single donor, thus making commonalities in recognition uncertain. Here we report the cocrystal structure of V1V2 with antibody CH03 from a second donor and model Env interactions of antibody CAP256-VRC26 from a third donor. These V1V2-directed bNAbs used strand-strand interactions between a protruding antibody loop and a V1V2 strand but differed in their N-glycan recognition. Ontogeny analysis indicated that protruding loops develop early, and glycan interactions mature over time. Altogether, the multidonor information suggested that V1V2-directed bNAbs form an 'extended class', for which we engineered ontogeny-specific antigens: Env trimers with chimeric V1V2s that interacted with inferred ancestor and intermediate antibodies. The ontogeny-based design of vaccine antigens described here may provide a general means for eliciting antibodies of a desired class.
Organizational Affiliation:
Vaccine Research Center, National Institute of Allergy and Infectious Diseases, National Institutes of Health (NIH), Bethesda, Maryland, USA.