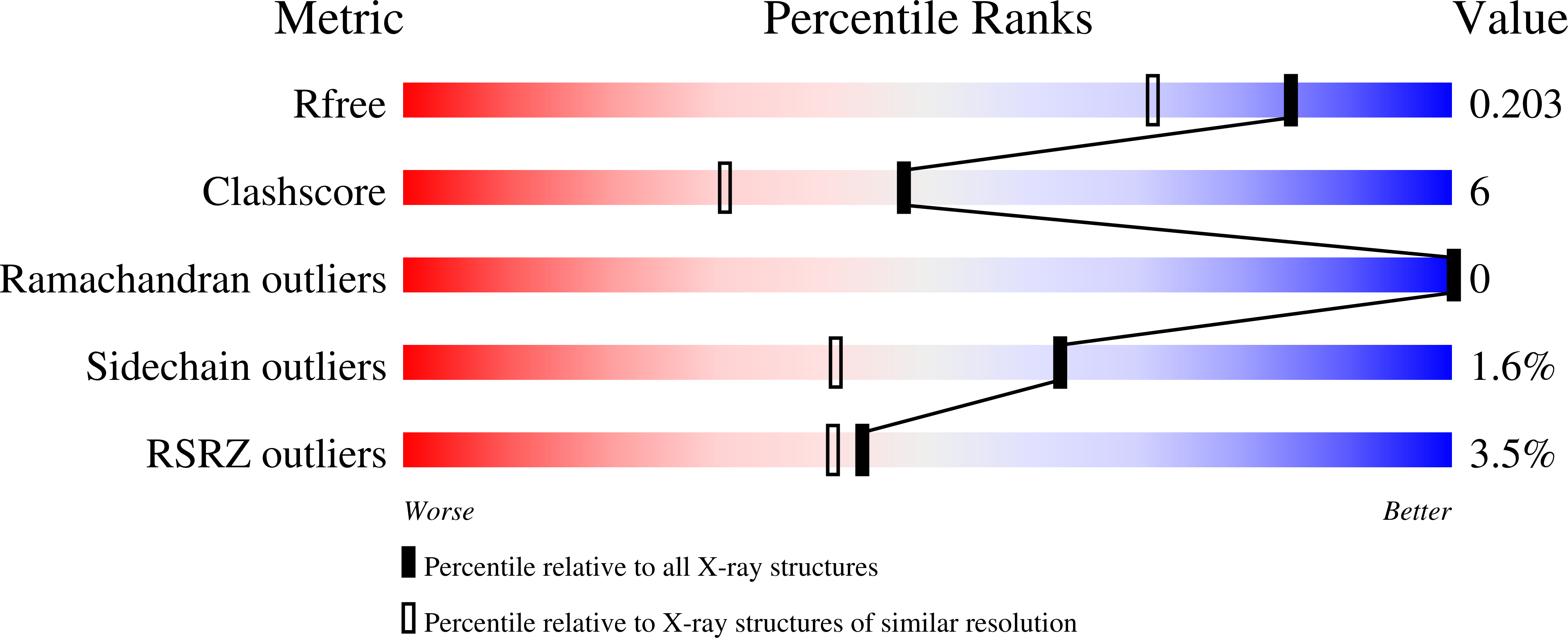
Crystal structure of amidohydrolase map2389c (target EFI-500390) from Mycobacterium avium
Patskovsky, Y., Toro, R., Bhosle, R., Hillerich, B., Seidel, R.D., Washington, E., Scott Glenn, A., Chowdhury, S., Evans, B., Hammonds, J., Zencheck, W.D., Imker, H.J., Gerlt, J.A., Raushel, F.M., Almo, S.C.To be published.