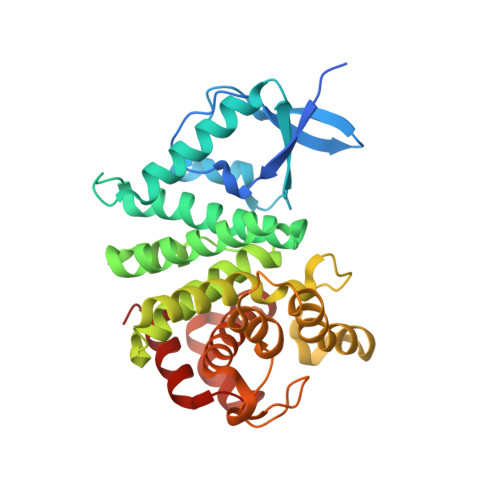
The structure of an unconventional HD-GYP protein from Bdellovibrio reveals the roles of conserved residues in this class of cyclic-di-GMP phosphodiesterases.
Lovering, A.L., Capeness, M.J., Lambert, C., Hobley, L., Sockett, R.E.(2011) mBio 2
- PubMed: 21990613
- DOI: https://doi.org/10.1128/mBio.00163-11
- Primary Citation of Related Structures:
3TM8, 3TMB, 3TMC, 3TMD - PubMed Abstract:
Cyclic-di-GMP is a near-ubiquitous bacterial second messenger that is important in localized signal transmission during the control of various processes, including virulence and switching between planktonic and biofilm-based lifestyles. Cyclic-di-GMP is synthesized by GGDEF diguanylate cyclases and hydrolyzed by EAL or HD-GYP phosphodiesterases, with each functional domain often appended to distinct sensory modules. HD-GYP domain proteins have resisted structural analysis, but here we present the first structural representative of this family (1.28 Å), obtained using the unusual Bd1817 HD-GYP protein from the predatory bacterium Bdellovibrio bacteriovorus. Bd1817 lacks the active-site tyrosine present in most HD-GYP family members yet remains an excellent model of their features, sharing 48% sequence similarity with the archetype RpfG. The protein structure is highly modular and thus provides a basis for delineating domain boundaries in other stimulus-dependent homologues. Conserved residues in the HD-GYP family cluster around a binuclear metal center, which is observed complexed to a molecule of phosphate, providing information on the mode of hydroxide ion attack on substrate. The fold and active site of the HD-GYP domain are different from those of EAL proteins, and restricted access to the active-site cleft is indicative of a different mode of activity regulation. The region encompassing the GYP motif has a novel conformation and is surface exposed and available for complexation with binding partners, including GGDEF proteins. It is becoming apparent that many bacteria use the signaling molecule cyclic-di-GMP to regulate a variety of processes, most notably, transitions between motility and sessility. Importantly, this regulation is central to several traits implicated in chronic disease (adhesion, biofilm formation, and virulence gene expression). The mechanisms of cyclic-di-GMP synthesis via GGDEF enzymes and hydrolysis via EAL enzymes have been suggested by the analysis of several crystal structures, but no information has been available to date for the unrelated HD-GYP class of hydrolases. Here we present the multidomain structure of an unusual member of the HD-GYP family from the predatory bacterium Bdellovibrio bacteriovorus and detail the features that distinguish it from the wider structural family of general HD fold hydrolases. The structure reveals how a binuclear iron center is formed from several conserved residues and provides a basis for understanding HD-GYP family sequence requirements for c-di-GMP hydrolysis.
- School of Biosciences, University of Birmingham, Birmingham, United Kingdom. a.lovering@bham.ac.uk
Organizational Affiliation: