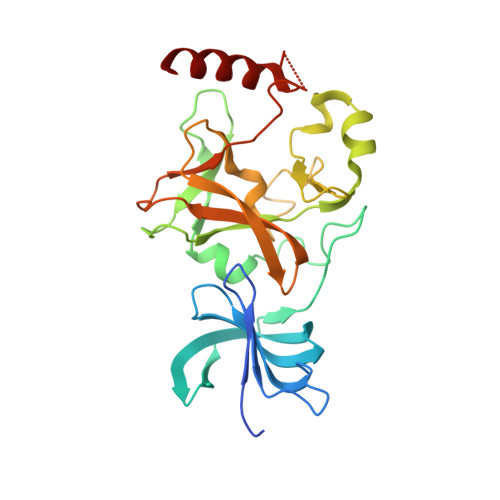
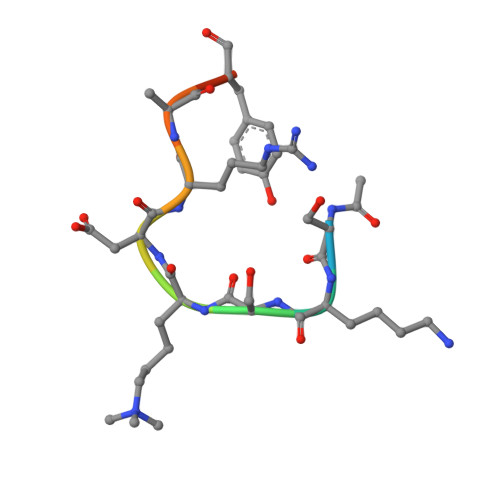
SET7/9 catalytic mutants reveal the role of active site water molecules in lysine multiple methylation.
Del Rizzo, P.A., Couture, J.F., Dirk, L.M., Strunk, B.S., Roiko, M.S., Brunzelle, J.S., Houtz, R.L., Trievel, R.C.(2010) J Biol Chem 285: 31849-31858
- PubMed: 20675860
- DOI: https://doi.org/10.1074/jbc.M110.114587
- Primary Citation of Related Structures:
3M53, 3M54, 3M55, 3M56, 3M57, 3M58, 3M59, 3M5A - PubMed Abstract:
SET domain lysine methyltransferases (KMTs) methylate specific lysine residues in histone and non-histone substrates. These enzymes also display product specificity by catalyzing distinct degrees of methylation of the lysine ε-amino group. To elucidate the molecular mechanism underlying this specificity, we have characterized the Y245A and Y305F mutants of the human KMT SET7/9 (also known as KMT7) that alter its product specificity from a monomethyltransferase to a di- and a trimethyltransferase, respectively. Crystal structures of these mutants in complex with peptides bearing unmodified, mono-, di-, and trimethylated lysines illustrate the roles of active site water molecules in aligning the lysine ε-amino group for methyl transfer with S-adenosylmethionine. Displacement or dissociation of these solvent molecules enlarges the diameter of the active site, accommodating the increasing size of the methylated ε-amino group during successive methyl transfer reactions. Together, these results furnish new insights into the roles of active site water molecules in modulating lysine multiple methylation by SET domain KMTs and provide the first molecular snapshots of the mono-, di-, and trimethyl transfer reactions catalyzed by these enzymes.
Organizational Affiliation:
Department of Biological Chemistry, University of Michigan, Ann Arbor, Michigan 48109, USA.