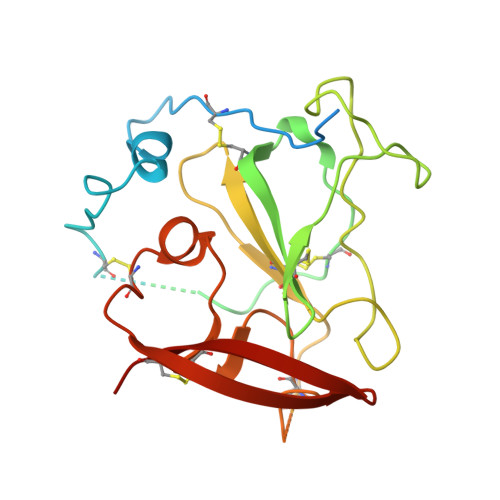
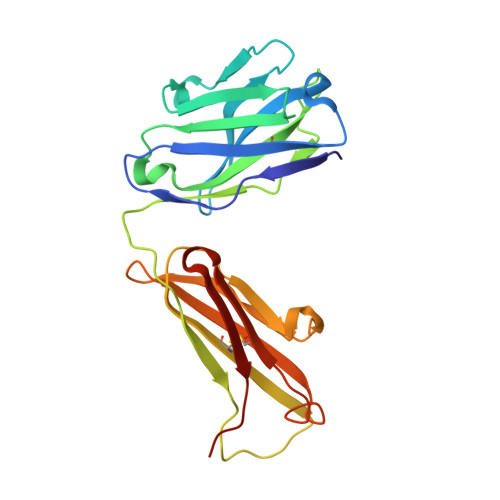
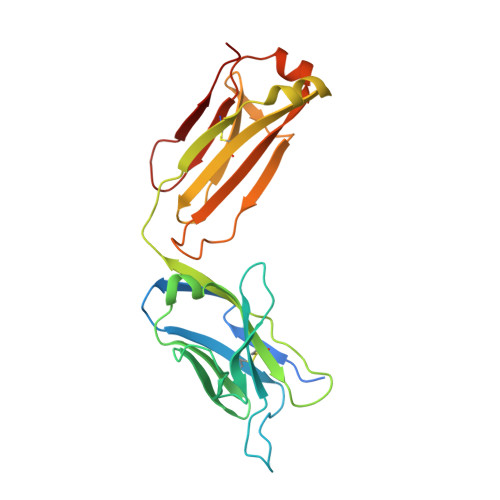
Structural insights into hepatitis C virus receptor binding and entry.
Kumar, A., Hossain, R.A., Yost, S.A., Bu, W., Wang, Y., Dearborn, A.D., Grakoui, A., Cohen, J.I., Marcotrigiano, J.(2021) Nature 598: 521-525
- PubMed: 34526719
- DOI: https://doi.org/10.1038/s41586-021-03913-5
- Primary Citation of Related Structures:
7MWS, 7MWW, 7MWX - PubMed Abstract:
Hepatitis C virus (HCV) infection is a causal agent of chronic liver disease, cirrhosis and hepatocellular carcinoma in humans, and afflicts more than 70 million people worldwide. The HCV envelope glycoproteins E1 and E2 are responsible for the binding of the virus to the host cell, but the exact entry process remains undetermined 1 . The majority of broadly neutralizing antibodies block interaction between HCV E2 and the large extracellular loop (LEL) of the cellular receptor CD81 (CD81-LEL) 2 . Here we show that low pH enhances the binding of CD81-LEL to E2, and we determine the crystal structure of E2 in complex with an antigen-binding fragment (2A12) and CD81-LEL (E2-2A12-CD81-LEL); E2 in complex with 2A12 (E2-2A12); and CD81-LEL alone. After binding CD81, residues 418-422 in E2 are displaced, which allows for the extension of an internal loop consisting of residues 520-539. Docking of the E2-CD81-LEL complex onto a membrane-embedded, full-length CD81 places the residues Tyr529 and Trp531 of E2 proximal to the membrane. Liposome flotation assays show that low pH and CD81-LEL increase the interaction of E2 with membranes, whereas structure-based mutants of Tyr529, Trp531 and Ile422 in the amino terminus of E2 abolish membrane binding. These data support a model in which acidification and receptor binding result in a conformational change in E2 in preparation for membrane fusion.
Organizational Affiliation:
Structural Virology Section, Laboratory of Infectious Diseases, National Institute of Allergy and Infectious Diseases, National Institutes of Health, Bethesda, MD, USA.