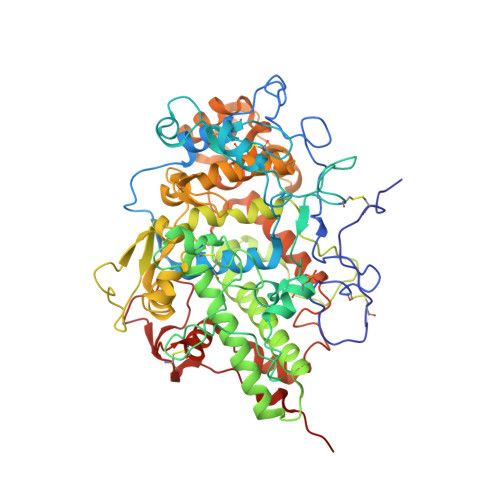
Design of anti-thyroid drugs: Binding studies and structure determination of the complex of lactoperoxidase with 2-mercaptoimidazole at 2.30 angstrom resolution
Sirohi, H.V., Singh, P.K., Iqbal, N., Sharma, P., Singh, A.K., Kaur, P., Sharma, S., Singh, T.P.(2017) Proteins 85: 1882-1890
- PubMed: 28653416
- DOI: https://doi.org/10.1002/prot.25342
- Primary Citation of Related Structures:
5GH0 - PubMed Abstract:
Lactoperoxidase (LPO) belongs to mammalian heme peroxidase superfamily, which also includes myeloperoxidase (MPO), eosinophil peroxidase (EPO), and thyroid peroxidase (TPO). LPO catalyzes the oxidation of a number of substrates including thiocyanate while TPO catalyzes the biosynthesis of thyroid hormones. LPO is also been shown to catalyze the biosynthesis of thyroid hormones indicating similar functional and structural properties. The binding studies showed that 2-mercaptoimidazole (MZY) bound to LPO with a dissociation constant of 0.63 µM. The inhibition studies showed that the value of IC 50 was 17 µM. The crystal structure of the complex of LPO with MZY showed that MZY bound to LPO in the substrate-binding site on the distal heme side. MZY was oriented in the substrate-binding site in such a way that the sulfur atom is at a distance of 2.58 Å from the heme iron. Previously, a similar compound, 3-amino-1,2,4-triazole (amitrole) was also shown to bind to LPO in the substrate-binding site on the distal heme side. The amino nitrogen atom of amitrole occupied the same position as that of sulfur atom in the present structure indicating a similar mode of binding. Recently, the structure of the complex of LPO with a potent antithyroid drug, 1-methylimidazole-2-thiol (methimazole, MMZ) was also determined. It showed that MMZ bound to LPO in the substrate-binding site on the distal heme side with 2 orientations. The position of methyl group was same in the 2 orientations while the positions of sulfur atom differed indicating a higher preference for a methyl group.
Organizational Affiliation:
Department of Biophysics, All India Institute of Medical Sciences, New Delhi.