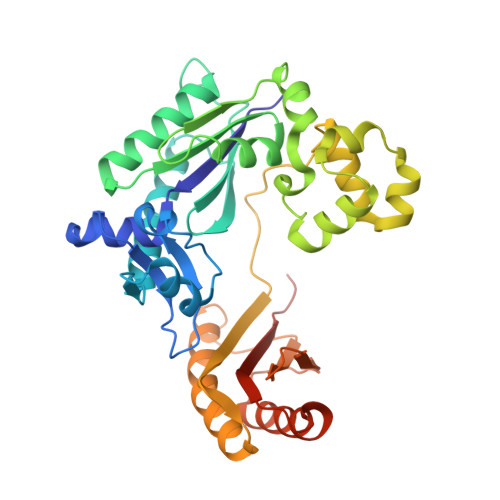
A strategically located serine residue is critical for the mutator activity of DNA polymerase IV from Escherichia coli.
Sharma, A., Kottur, J., Narayanan, N., Nair, D.T.(2013) Nucleic Acids Res 41: 5104-5114
- PubMed: 23525461
- DOI: https://doi.org/10.1093/nar/gkt146
- Primary Citation of Related Structures:
4IR1, 4IR9, 4IRC, 4IRD, 4IRK - PubMed Abstract:
The Y-family DNA polymerase IV or PolIV (Escherichia coli) is the founding member of the DinB family and is known to play an important role in stress-induced mutagenesis. We have determined four crystal structures of this enzyme in its pre-catalytic state in complex with substrate DNA presenting the four possible template nucleotides that are paired with the corresponding incoming nucleotide triphosphates. In all four structures, the Ser42 residue in the active site forms interactions with the base moieties of the incipient Watson-Crick base pair. This residue is located close to the centre of the nascent base pair towards the minor groove. In vitro and in vivo assays show that the fidelity of the PolIV enzyme increases drastically when this Ser residue was mutated to Ala. In addition, the structure of PolIV with the mismatch A:C in the active site shows that the Ser42 residue plays an important role in stabilizing dCTP in a conformation compatible with catalysis. Overall, the structural, biochemical and functional data presented here show that the Ser42 residue is present at a strategic location to stabilize mismatches in the PolIV active site, and thus facilitate the appearance of transition and transversion mutations.
Organizational Affiliation:
National Centre for Biological Sciences (NCBS-TIFR), UAS-GKVK Campus, Bellary Road, Bangalore 560065, India.