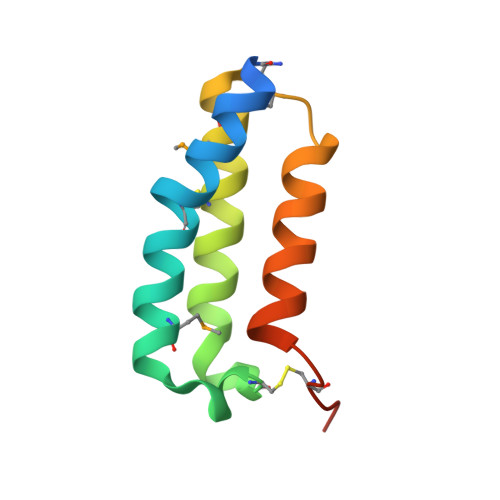
Crystal structure of the human receptor activity-modifying protein 1 extracellular domain.
Kusano, S., Kukimoto-Niino, M., Akasaka, R., Toyama, M., Terada, T., Shirouzu, M., Shindo, T., Yokoyama, S.(2008) Protein Sci 17: 1907-1914
- PubMed: 18725456
- DOI: https://doi.org/10.1110/ps.036012.108
- Primary Citation of Related Structures:
2YX8 - PubMed Abstract:
Receptor activity-modifying protein (RAMP) 1 forms a heterodimer with calcitonin receptor-like receptor (CRLR) and regulates its transport to the cell surface. The CRLR.RAMP1 heterodimer functions as a specific receptor for calcitonin gene-related peptide (CGRP). Here, we report the crystal structure of the human RAMP1 extracellular domain. The RAMP1 structure is a three-helix bundle that is stabilized by three disulfide bonds. The RAMP1 residues important for cell-surface expression of the CRLR.RAMP1 heterodimer are clustered to form a hydrophobic patch on the molecular surface. The hydrophobic patch is located near the tryptophan residue essential for binding of the CGRP antagonist, BIBN4096BS. These results suggest that the hydrophobic patch participates in the interaction with CRLR and the formation of the ligand-binding pocket when it forms the CRLR.RAMP1 heterodimer.
Organizational Affiliation:
RIKEN Systems and Structural Biology Center, Yokohama 230-0045, Japan.