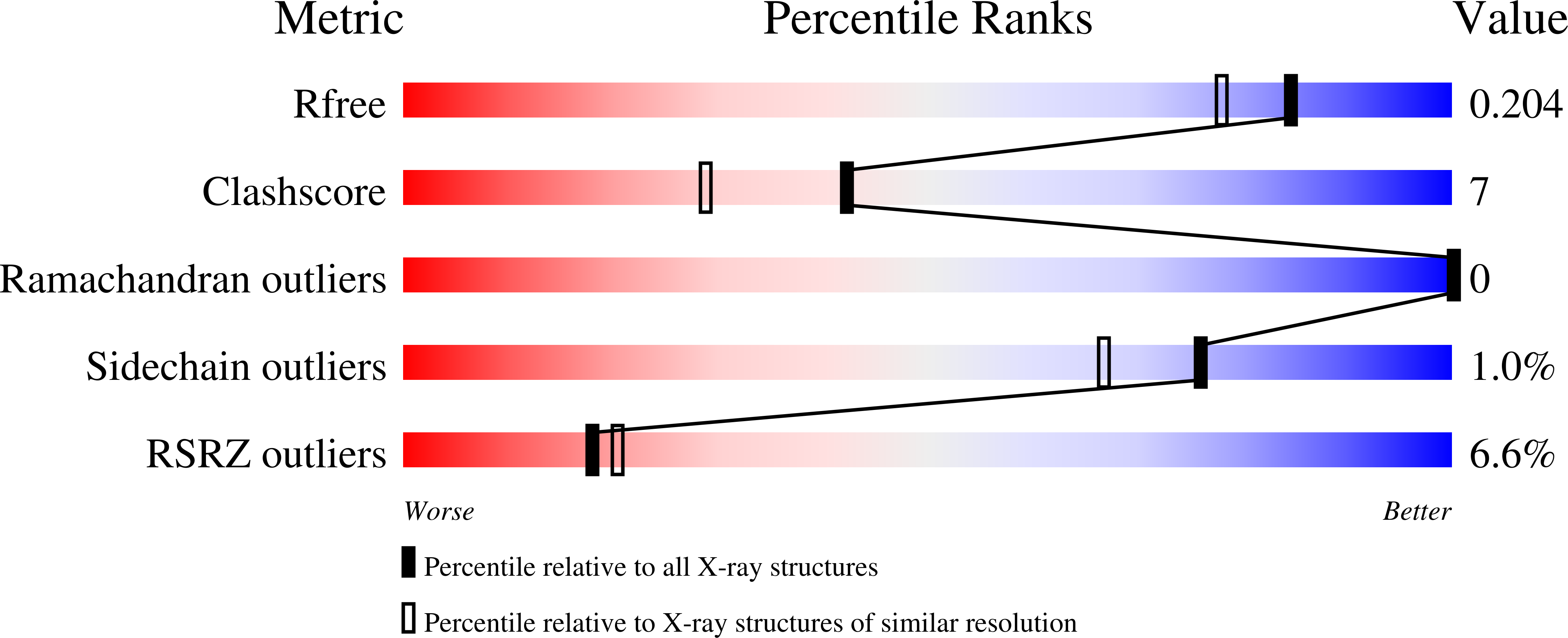
Agrobacterium tumefaciens VirC2 enhances T-DNA transfer and virulence through its C-terminal ribbon-helix-helix DNA-binding fold
Lu, J., den Dulk-Ras, A., Hooykaas, P.J., Glover, J.N.(2009) Proc Natl Acad Sci U S A 106: 9643-9648
- PubMed: 19482939
- DOI: https://doi.org/10.1073/pnas.0812199106
- Primary Citation of Related Structures:
2RH3 - PubMed Abstract:
Agrobacterium tumefaciens VirC2 stimulates processing of single-stranded T-DNA that is translocated into plants to induce tumor formation, but how VirC2 functions is unclear. Here, we report the 1.7-A X-ray crystal structure of its trypsin-resistant C-terminal domain, VirC2(82-202), which reveals a form of the ribbon-helix-helix (RHH) DNA-binding fold contained within a single polypeptide chain. DNA-binding assays and mutagenesis indicate that VirC2 uses this RHH fold to bind double-stranded DNA but not single-stranded DNA. Mutations that severely affect VirC2 DNA binding are highly deleterious for both T-DNA transfer into yeast and the virulence of A. tumefaciens in different plants including Nicotiana glauca and Kalanchoe daigremontiana. These data suggest that VirC2 enhances T-DNA transfer and virulence through DNA binding with its RHH fold. The RHH fold of VirC2 is the first crystal structure representing a group of predicted RHH proteins that facilitate endonucleolytic processing of DNA for horizontal gene transfer.
Organizational Affiliation:
Department of Biochemistry, University of Alberta, Edmonton, Alberta, Canada T6G 2H7.