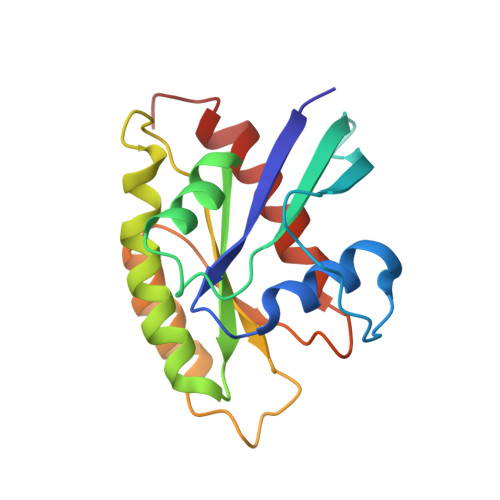
Solution structure and dynamics of the small GTPase RalB in its active conformation: significance for effector protein binding
Fenwick, R.B., Prasannan, S., Campbell, L.J., Nietlispach, D., Evetts, K.A., Camonis, J., Mott, H.R., Owen, D.(2009) Biochemistry 48: 2192-2206
- PubMed: 19166349
- DOI: https://doi.org/10.1021/bi802129d
- Primary Citation of Related Structures:
2KE5 - PubMed Abstract:
The small G proteins RalA/B have a crucial function in the regulatory network that couples extracellular signals with appropriate cellular responses. RalA/B are an important component of the Ras signaling pathway and, in addition to their role in membrane trafficking, are implicated in the initiation and maintenance of tumorigenic transformation of human cells. RalA and RalB share 85% sequence identity and collaborate in supporting cancer cell proliferation but have markedly different effects. RalA is important in mediating proliferation, while depletion of RalB results in transformed cells undergoing apoptosis. Crystal structures of RalA in the free form and in complex with its effectors, Sec5 and Exo84, have been solved. Here we have determined the solution structure of free RalB bound to the GTP analogue GMPPNP to an RMSD of 0.6 A. We show that, while the overall architecture of RalB is very similar to the crystal structure of RalA, differences exist in the switch regions, which are sensitive to the bound nucleotide. Backbone 15N dynamics suggest that there are four regions of disorder in RalB: the P-loop, switch I, switch II, and the loop comprising residues 116-121, which has a single residue insertion compared to RalA. 31P NMR data and the structure of RalB.GMPPNP show that the switch regions predominantly adopt state 1 (Ras nomenclature) in the unbound form, which in Ras is not competent to bind effectors. In contrast, 31P NMR analysis of RalB.GTP reveals that conformations corresponding to states 1 and 2 are both sampled in solution and that addition of an effector protein only partially stabilizes state 2.
- Department of Biochemistry, University of Cambridge, UK.
Organizational Affiliation: