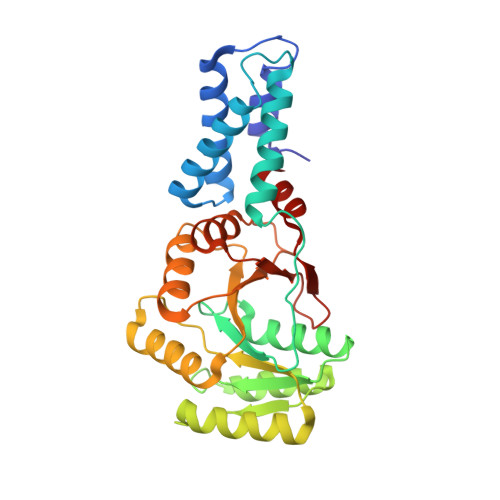
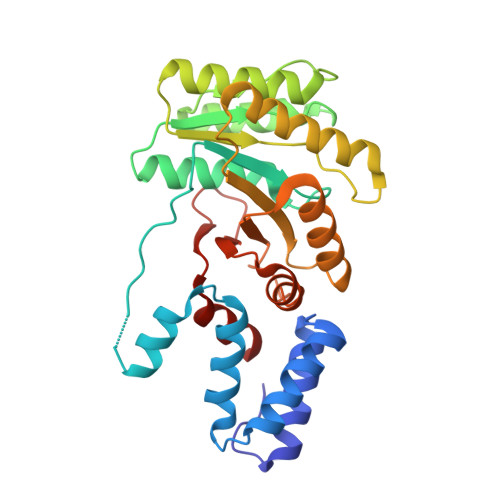
Structure of a Gdp:Alf(4) Complex of the Srp Gtpases Ffh and Ftsy, and Identification of a Peripheral Nucleotide Interaction Site.
Focia, P.J., Gawronski-Salerno, J., Coon V, J.S., Freymann, D.M.(2006) J Mol Biol 360: 631
- PubMed: 16780874
- DOI: https://doi.org/10.1016/j.jmb.2006.05.031
- Primary Citation of Related Structures:
2CNW - PubMed Abstract:
The signal recognition particle (SRP) GTPases Ffh and FtsY play a central role in co-translational targeting of proteins, assembling in a GTP-dependent manner to generate the SRP targeting complex at the membrane. A suite of residues in FtsY have been identified that are essential for the hydrolysis of GTP that accompanies disengagement. We have argued previously on structural grounds that this region mediates interactions that serve to activate the complex for disengagement and term it the activation region. We report here the structure of a complex of the SRP GTPases formed in the presence of GDP:AlF4. This complex accommodates the putative transition-state analog without undergoing significant change from the structure of the ground-state complex formed in the presence of the GTP analog GMPPCP. However, small shifts that do occur within the shared catalytic chamber may be functionally important. Remarkably, an external nucleotide interaction site was identified at the activation region, revealed by an unexpected contaminating GMP molecule bound adjacent to the catalytic chamber. This site exhibits conserved sequence and structural features that suggest a direct interaction with RNA plays a role in regulating the activity of the SRP targeting complex.
Organizational Affiliation:
Department of Molecular Pharmacology and Biological Chemistry, Feinberg School of Medicine, Northwestern University, 303 E. Chicago Ave., Chicago, IL 60611, USA. focia@northwestern.edu