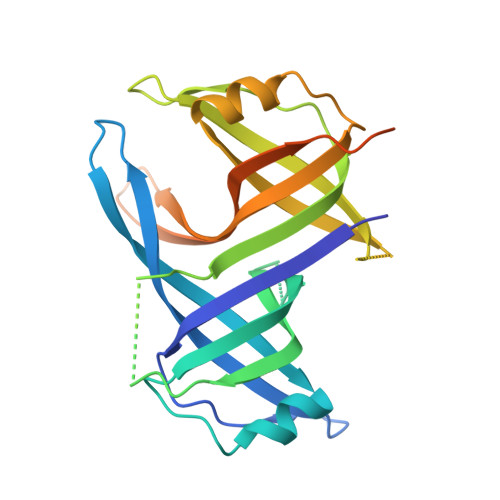
Structure of the single-stranded DNA-binding protein SSB from Thermus aquaticus.
Jedrzejczak, R., Dauter, M., Dauter, Z., Olszewski, M., Kur, J.(2006) Acta Crystallogr D Biol Crystallogr 62: 1407-1412
- PubMed: 17057346
- DOI: https://doi.org/10.1107/S0907444906036031
- Primary Citation of Related Structures:
2FXQ - PubMed Abstract:
The crystal structure of the single-stranded DNA-binding protein from Thermus aquaticus has been solved and refined at 1.85 A resolution. Two monomers, each encompassing two oligonucleotide/oligosaccharide-binding (OB) domains and a number of flexible beta-hairpin loops, form an oligomer of approximate D(2) symmetry typical of bacterial SSBs. Comparison with other SSB structures confirms considerable variability in the mode of oligomerization and aggregation of SSB oligomers.
Organizational Affiliation:
Synchrotron Radiation Research Section, MCL, National Cancer Institute, Argonne National Laboratory, Argonne, IL 60439, USA.