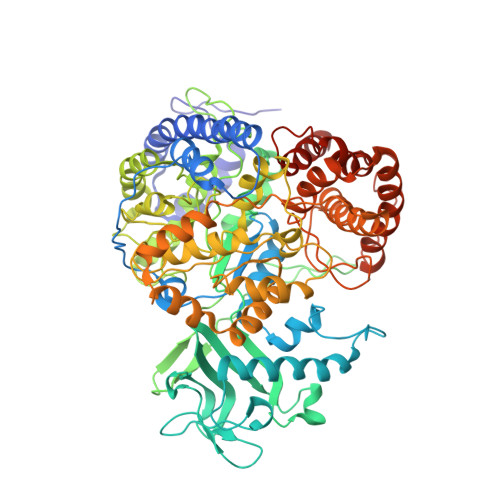
Solution NMR-derived global fold of a monomeric 82-kDa enzyme.
Tugarinov, V., Choy, W.-Y., Orekhov, V.Y., Kay, L.E.(2005) Proc Natl Acad Sci U S A 102: 622-627
- PubMed: 15637152
- DOI: https://doi.org/10.1073/pnas.0407792102
- Primary Citation of Related Structures:
1Y8B - PubMed Abstract:
The size of proteins that can be studied by solution NMR spectroscopy has increased significantly because of recent developments in methodology. Important experiments include those that make use of approaches that increase the lifetimes of NMR signals or that define the orientation of internuclear bond vectors with respect to a common molecular frame. The advances in NMR techniques are strongly coupled to isotope labeling methods that increase sensitivity and reduce the complexity of NMR spectra. We show that these developments can be exploited in structural studies of high-molecular-weight, single-polypeptide proteins, and we present the solution global fold of the monomeric 723-residue (82-kDa) enzyme malate synthase G from Escherichia coli, which has been extensively characterized by NMR in the past several years.
Organizational Affiliation:
Protein Engineering Network Centres of Excellence and Department of Medical Genetics, University of Toronto, Toronto, ON, Canada M5S 1A8.