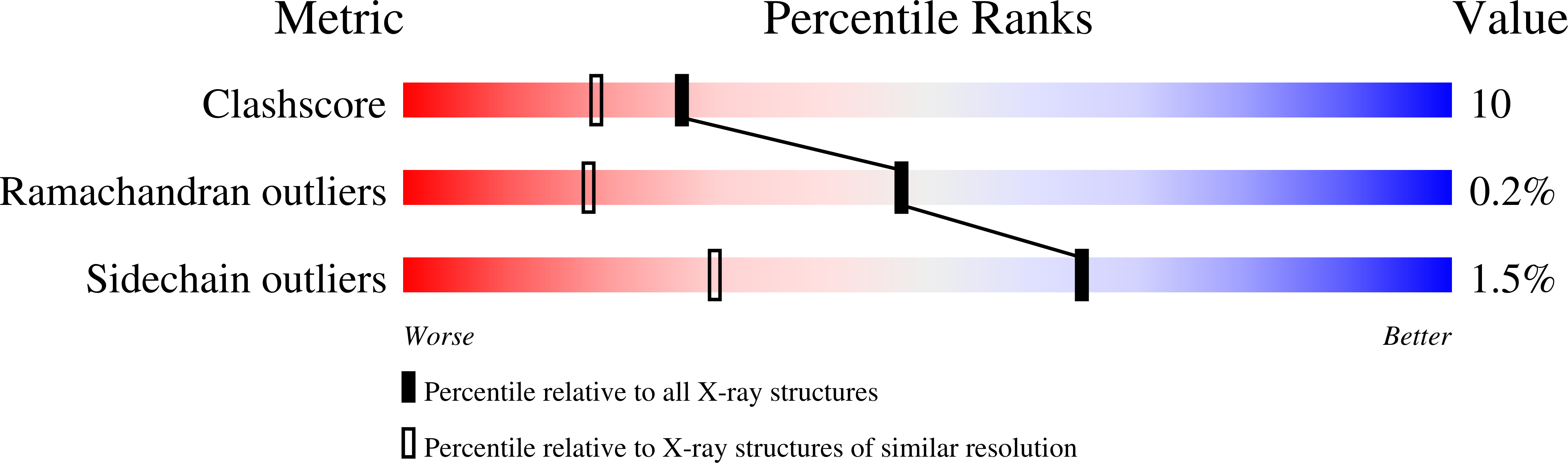Sub-atomic resolution crystal structure of cholesterol oxidase: What atomic resolution crystallography reveals about enzyme mechanism and the role of FAD cofactor in redox activity
Lario, P.I., Sampson, N., Vrielink, A.(2003) J Mol Biol 326: 1635-1650
- PubMed: 12595270
- DOI: https://doi.org/10.1016/s0022-2836(03)00054-8
- Primary Citation of Related Structures:
1MXT - PubMed Abstract:
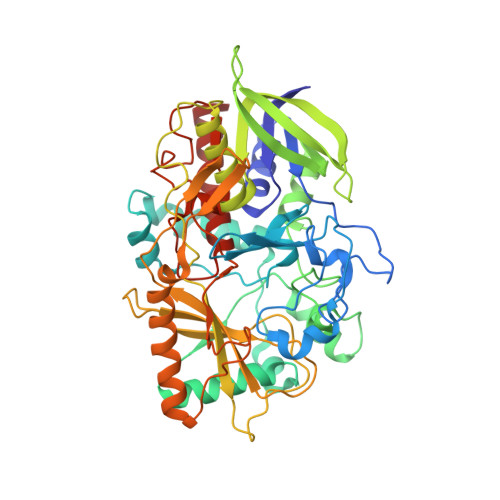
The crystal structure of cholesterol oxidase, a 56kDa flavoenzyme was anisotropically refined to 0.95A resolution. The final crystallographic R-factor and R(free) value is 11.0% and 13.2%, respectively. The quality of the electron density maps has enabled modeling of alternate conformations for 83 residues in the enzyme, many of which are located in the active site. The additional observed structural features were not apparent in the previous high-resolution structure (1.5A resolution) and have enabled the identification of a narrow tunnel leading directly to the isoalloxazine portion of the FAD prosthetic group. The hydrophobic nature of this narrow tunnel suggests it is the pathway for molecular oxygen to access the isoalloxazine group for the oxidative half reaction. Resolving the alternate conformations in the active site residues provides a model for the dynamics of substrate binding and a potential oxidation triggered gating mechanism involving access to the hydrophobic tunnel. This structure reveals that the NE2 atom of the active site histidine residue, H447, critical to the redox activity of this flavin oxidase, acts as a hydrogen bond donor rather than as hydrogen acceptor. The atomic resolution structure of cholesterol oxidase has revealed the presence of hydrogen atoms, dynamic aspects of the protein and how side-chain conformations are correlated with novel structural features such as the oxygen tunnel. This new structural information has provided us with the opportunity to re-analyze the roles played by specific residues in the mechanism of the enzyme.
Organizational Affiliation:
Department of Molecular, Cellular and Developmental Biology, Sinsheimer Laboratory, University of California Santa Cruz, 1156 High Street, Santa Cruz, CA 95064, USA.