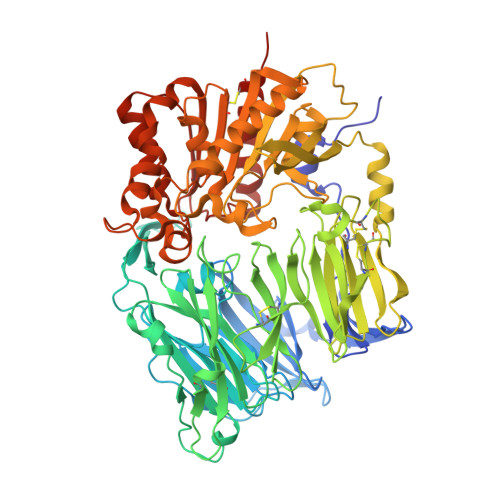
The structure and function of human dipeptidyl peptidase IV, possessing a unique eight-bladed beta-propeller fold.
Hiramatsu, H., Kyono, K., Higashiyama, Y., Fukushima, C., Shima, H., Sugiyama, S., Inaka, K., Yamamoto, A., Shimizu, R.(2003) Biochem Biophys Res Commun 302: 849-854
- PubMed: 12646248
- DOI: https://doi.org/10.1016/s0006-291x(03)00258-4
- Primary Citation of Related Structures:
1J2E - PubMed Abstract:
Dipeptidyl peptidase IV (DPPIV) is a serine protease, a member of the prolyl oligopeptidase (POP) family, and has been implicated in several diseases. Therefore, the development of DPPIV selective inhibitors, which are able to control the biological function of DPPIV, is important. We determined the crystal structure of human DPPIV at 2.6A resolution. The molecule consists of a unique eight-bladed beta-propeller domain in the N-terminal region and a serine protease domain in the C-terminal region. Also, the large "cave" structure, which is thought to control the access of the substrate, is found on the side of the beta-propeller fold. Comparison of the overall amino acid sequence between human DPPIV and POP shows low homology (12.9%). In this paper, we report the structure of human DPPIV, especially focusing on a unique eight-bladed beta-propeller domain. We also discuss the way for the access of the substrate to this domain.
Organizational Affiliation:
Discovery Research Laboratory, Tanabe Seiyaku Co. Ltd., 3-16-86 Kashima, Yodogawa-ku, Osaka 532-8505, Japan.